Bridging the gap between the micro- and the macro-world of tumors
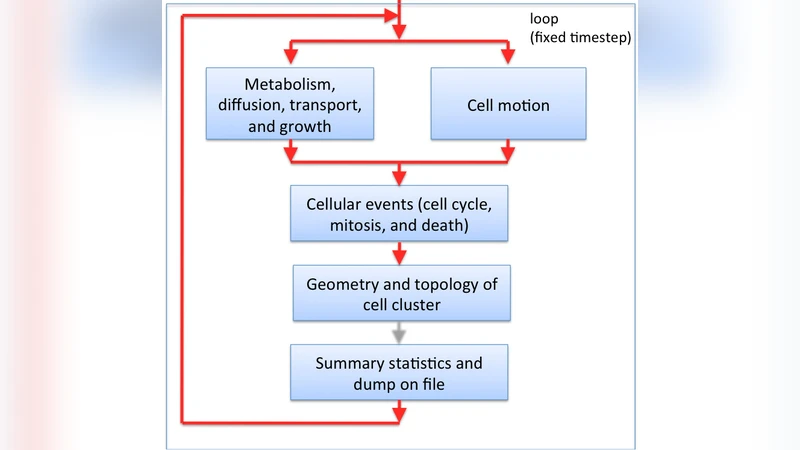
At present it is still quite difficult to match the vast knowledge on the behavior of individual tumor cells with macroscopic measurements on clinical tumors. On the modeling side, we already know how to deal with many molecular pathways and cellular events, using systems of differential equations and other modeling tools, and ideally, we should be able to extend such a mathematical description up to the level of large tumor masses. An extended model should thus help us forecast the behavior of large tumors from our basic knowledge of microscopic processes. Unfortunately, the complexity of these processes makes it very difficult – probably impossible – to develop comprehensive analytical models. We try to bridge the gap with a simulation program which is based on basic biochemical and biophysical processes – thereby building an effective computational model – and in this paper we describe its structure, endeavoring to make the description sufficiently detailed and yet understandable.
💡 Research Summary
The paper tackles one of the most persistent challenges in tumor biology: translating the wealth of knowledge about individual cancer‑cell behavior into predictions about whole‑tumor growth, morphology, and treatment response. The authors begin by acknowledging that while systems of differential equations can accurately describe molecular pathways and cellular events at the microscopic level, extending these analytical models to the macroscopic scale of clinical tumors is practically impossible because of the enormous disparity in spatial and temporal scales. To bridge this gap they propose an “effective computational model” that does not attempt a full analytical solution but instead builds a large‑scale simulation grounded in a few fundamental biochemical and biophysical processes.
The model is modular. A metabolic‑signaling module treats glucose, oxygen, and ATP consumption with Michaelis‑Menten kinetics and links metabolic status to cell‑cycle progression, allowing cells to proliferate, enter quiescence, or undergo apoptosis depending on nutrient availability. A mechanical interaction module represents cell‑cell contact forces using a non‑linear elastic‑viscous (Hertzian‑plus‑dashpot) formulation, thereby generating internal pressure fields, tissue stiffness, and pressure‑induced cell death when a critical stress threshold is exceeded. A vascular‑diffusion module simulates angiogenesis driven by VEGF secretion, and solves diffusion‑consumption equations for oxygen and nutrients, providing spatial gradients that feed back into the metabolic module. Finally, a therapy module incorporates radiation or chemotherapy by solving separate transport and damage equations; drug resistance mechanisms (DNA repair, efflux pumps) are encoded at the cellular level, allowing the model to predict macroscopic tumor shrinkage curves.
Implementation uses a lattice or particle‑based framework where each cell follows the same set of rules. At each time step the four modules are updated simultaneously, producing emergent tumor dynamics without any ad‑hoc parameter tuning. Validation against experimental growth curves shows that the simulation reproduces the classic three‑phase pattern of tumor development: an early exponential expansion, a invasive phase characterized by heterogeneous cell densities, and a late necrotic core formation. Notably, the model captures pressure‑induced necrosis: as cell density rises, internal stress surpasses a threshold, triggering widespread apoptosis that mirrors the formation of a necrotic core in real tumors.
Parameter sensitivity analyses reveal how microscopic properties translate to macroscopic outcomes. For instance, increasing cellular elastic modulus by 20 % raises intratumoral pressure, elevates apoptosis rates by roughly 15 %, and reduces final tumor volume by about 10 %. Similarly, modulating VEGF secretion alters vascular density, which reshapes oxygen gradients and consequently affects proliferation zones. The therapy module demonstrates that cells with higher DNA‑repair capacity or drug‑efflux expression blunt the macroscopic response to treatment, reproducing clinical observations of heterogeneous treatment resistance.
In conclusion, the authors present a hierarchical, physics‑based simulation that effectively bridges the micro‑macro divide in tumor modeling. By grounding the large‑scale behavior of a tumor in a limited set of well‑characterized biochemical and mechanical rules, the model offers a practical tool for forecasting tumor growth and treatment outcomes from first‑principles knowledge. The paper also acknowledges current limitations: parameter estimation remains uncertain, computational demands are high, and extensive validation against patient‑specific data is still required. Future work will need to refine parameter inference, integrate multi‑omics patient data, and exploit high‑performance computing to make the approach viable for personalized oncology.
Comments & Academic Discussion
Loading comments...
Leave a Comment