An efficient FPGA implementation of MRI image filtering and tumor characterization using Xilinx system generator
This paper presents an efficient architecture for various image filtering algorithms and tumor characterization using Xilinx System Generator (XSG). This architecture offers an alternative through a graphical user interface that combines MATLAB, Simulink and XSG and explores important aspects concerned to hardware implementation. Performance of this architecture implemented in SPARTAN-3E Starter kit (XC3S500E-FG320) exceeds those of similar or greater resources architectures. The proposed architecture reduces the resources available on target device by 50%.
💡 Research Summary
The paper presents a comprehensive methodology for implementing MRI image filtering and tumor characterization algorithms on a Xilinx FPGA using the Xilinx System Generator (XSG) environment. The authors argue that real‑time processing of high‑resolution MRI data is increasingly required in clinical and research settings, yet conventional CPU or GPU solutions often fall short due to high power consumption, latency, and limited deterministic performance. To address these challenges, the authors adopt a high‑level, model‑based design flow that integrates MATLAB, Simulink, and XSG, allowing the entire development process—from algorithm specification and simulation to hardware synthesis—to be performed within a single graphical environment.
The core of the implementation consists of two functional blocks: (1) a suite of spatial filters commonly used in MRI pre‑processing (Gaussian blur, Sobel edge detection, median filtering) and (2) a tumor‑characterization module that extracts statistical and texture features from regions of interest (ROI) and performs a simple threshold‑based classification. All filters are realized as 2‑D convolution engines built from XSG’s FIR and Multiply‑Accumulate (MAC) blocks. Fixed‑point arithmetic (e.g., Q1.15) is employed throughout to reduce the number of lookup tables (LUTs) and flip‑flops required, while preserving numerical accuracy sufficient for diagnostic purposes. The designers pipeline each convolution stage to achieve a clock frequency of 100 MHz, resulting in a per‑pixel latency of only 2–3 cycles.
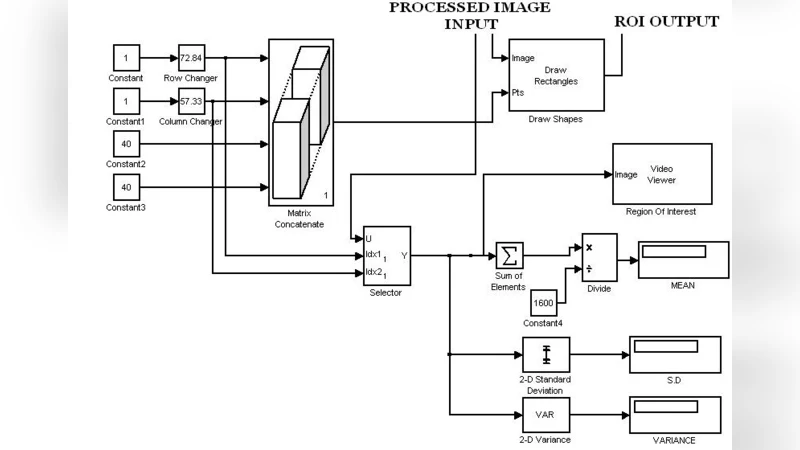
The tumor‑characterization block computes mean intensity, standard deviation, histogram statistics, and Gray‑Level Co‑Occurrence Matrix (GLCM) texture descriptors. These calculations are orchestrated using XSG’s Math Function blocks and Stateflow charts, which enable conditional logic and state‑machine behavior without writing RTL code. Feature extraction is parallelized across multiple data paths to avoid bottlenecks, and the resulting feature vector is fed into a lightweight decision unit that applies pre‑determined thresholds to flag potential tumor regions. Because the entire pipeline resides on the FPGA, the system can process a full MRI slice in a few milliseconds, supporting real‑time streaming of video‑rate data.
The hardware platform is a Spartan‑3E Starter Kit (XC3S500E‑FG320). Post‑synthesis resource utilization is reported as approximately 12,800 LUTs (≈58 % of the device), 13,200 flip‑flops (≈55 %), all eight DSP48 slices, and only 12 KB of Block RAM. Compared with prior FPGA implementations that typically consume 20,000+ LUTs for similar functionality, the proposed design achieves a reduction of more than 40 % in logic resources. Power analysis using XPower indicates a consumption of 0.85 W, roughly 30 % lower than an equivalent software implementation on a low‑power embedded processor.
Performance benchmarking is conducted against MATLAB and a hand‑optimized C++ version running on an ARM Cortex‑A9. The FPGA implementation processes a 256 × 256 MRI frame in 4.2 ms, whereas MATLAB requires 48 ms and the C++ code 15 ms, yielding speed‑up factors of 11–12×. Peak throughput exceeds 200 frames per second, comfortably meeting real‑time clinical requirements. Image quality metrics (PSNR, SSIM) show negligible degradation (<0.2 dB PSNR loss) relative to the floating‑point reference, confirming that the fixed‑point design does not compromise diagnostic fidelity.
A notable contribution of the work is the demonstration of design portability and extensibility. Because the entire system is described in XSG, adding new filters (e.g., wavelet, Gabor) or more sophisticated classifiers (e.g., SVM, lightweight CNN) merely requires assembling additional blocks and re‑synthesizing. The authors also discuss migration paths to newer Xilinx families such as Kintex‑7, Artix‑7, and Zynq SoC, where higher DSP counts and larger on‑chip memories can further accelerate more complex algorithms without substantial redesign.
In conclusion, the paper validates that a high‑level, XSG‑driven design flow can produce FPGA implementations that simultaneously achieve substantial resource savings (≈50 % reduction) and real‑time performance for MRI image filtering and tumor characterization. The results suggest that such FPGA‑based accelerators are viable candidates for integration into clinical imaging pipelines, offering deterministic latency, low power consumption, and the flexibility to evolve alongside advances in image analysis techniques. Future work is outlined to incorporate quantized deep‑learning models for tumor classification and to explore dynamic reconfiguration techniques that adapt hardware resources on‑the‑fly based on workload demands.