Structure Learning of Probabilistic Graphical Models: A Comprehensive Survey
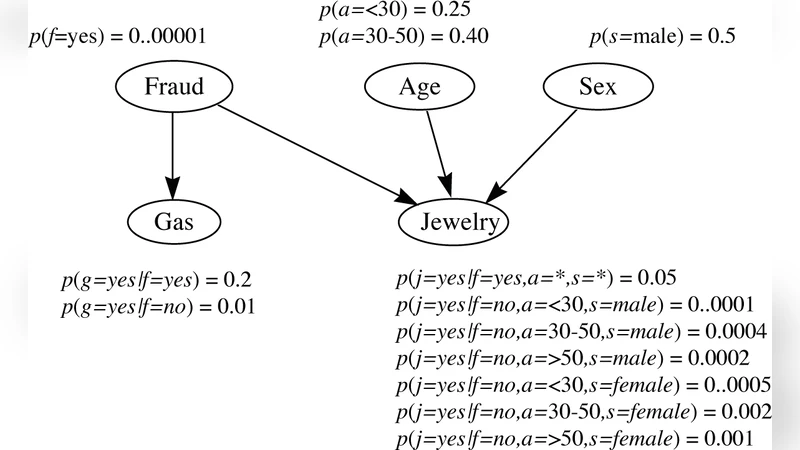
Probabilistic graphical models combine the graph theory and probability theory to give a multivariate statistical modeling. They provide a unified description of uncertainty using probability and complexity using the graphical model. Especially, graphical models provide the following several useful properties: - Graphical models provide a simple and intuitive interpretation of the structures of probabilistic models. On the other hand, they can be used to design and motivate new models. - Graphical models provide additional insights into the properties of the model, including the conditional independence properties. - Complex computations which are required to perform inference and learning in sophisticated models can be expressed in terms of graphical manipulations, in which the underlying mathematical expressions are carried along implicitly. The graphical models have been applied to a large number of fields, including bioinformatics, social science, control theory, image processing, marketing analysis, among others. However, structure learning for graphical models remains an open challenge, since one must cope with a combinatorial search over the space of all possible structures. In this paper, we present a comprehensive survey of the existing structure learning algorithms.
💡 Research Summary
This survey provides a comprehensive overview of structure learning methods for probabilistic graphical models (PGMs), covering both undirected models such as Markov Random Fields (MRFs) and Gaussian Graphical Models (GGMs), and directed models exemplified by Bayesian networks. After a concise introduction to the fundamental concepts—graphs, random variables, Markov blankets, d‑separation, and factorization—the paper categorizes existing algorithms into four major families: constraint‑based, score‑based, regression‑based, and hybrid/other approaches.
Constraint‑based methods (e.g., SGS, PC, GS) rely on conditional independence tests to prune or add edges. They are attractive because they reduce the search space dramatically, but their performance is highly sensitive to statistical test errors, especially in high‑dimensional, low‑sample regimes, and the number of tests grows combinatorially with the number of variables.
Score‑based techniques assign a numerical quality measure to each candidate graph. The survey details three widely used scores—Minimum Description Length (MDL), Bayesian Dirichlet equivalent (BDe), and Bayesian Information Criterion (BIC)—explaining how each balances model fit against complexity. Optimization strategies range from exhaustive search over the full DAG space to more tractable order‑based searches, greedy hill‑climbing, dynamic programming, and Markov chain Monte Carlo (MCMC) sampling. While score‑based methods can, in principle, locate globally optimal structures, they require sufficient statistics, appropriate priors, and still face exponential search complexity.
Regression‑based algorithms treat each variable as a response and regress it on a subset of other variables, using techniques such as Lasso, Elastic Net, or likelihood‑based objectives. In the Gaussian case, sparsity in the precision matrix directly corresponds to the graph’s edge set, making penalized likelihood a natural choice. These methods excel in high‑dimensional settings with limited data, but they are less suited for discrete or highly non‑linear relationships unless extended with kernel or generalized linear models.
Hybrid and other approaches combine the strengths of the previous families or introduce entirely different paradigms. Examples include: (i) initializing a structure with a clustering or community‑detection step and then refining it via constraint‑based tests; (ii) employing Boolean satisfiability solvers for discrete networks; (iii) leveraging information‑theoretic criteria such as mutual information or Minimum Description Length directly on data; and (iv) using matrix factorization (e.g., NMF, PCA) to uncover latent factors that define inter‑node connections. These methods often target specific application domains—gene‑regulatory networks, social networks, image segmentation—where domain knowledge can guide the algorithmic design.
The paper systematically compares each algorithm in terms of computational complexity, sample complexity, underlying assumptions (e.g., linearity, Gaussianity, faithfulness), and typical use cases. Tables summarizing these trade‑offs are provided, along with illustrative case studies from bioinformatics, computer vision, and social science.
Finally, the authors discuss open challenges: structure learning remains NP‑hard, making exact solutions infeasible for large graphs; statistical tests can be unreliable under noise or missing data; score functions may be misspecified; and scalability to massive, streaming datasets is still limited. Emerging research directions highlighted include scalable approximations via distributed computing, Bayesian non‑parametric priors (Dirichlet processes, Indian buffet processes) that allow the number of edges to grow with data, robust independence testing, and exploiting known network topologies such as small‑world or scale‑free properties.
In summary, this survey consolidates the state‑of‑the‑art in PGM structure learning, offering a clear taxonomy, detailed methodological insights, and practical guidance for researchers aiming to select or develop algorithms suited to their specific data and domain constraints.
Comments & Academic Discussion
Loading comments...
Leave a Comment