Tractability results for the Double-Cut-and-Join circular median problem
The circular median problem in the Double-Cut-and-Join (DCJ) distance asks to find, for three given genomes, a fourth circular genome that minimizes the sum of the mutual distances with the three other ones. This problem has been shown to be NP-complete. We show here that, if the number of vertices of degree 3 in the breakpoint graph of the three input genomes is fixed, then the problem is tractable
💡 Research Summary
**
The paper addresses the circular median problem under the Double‑Cut‑and‑Join (DCJ) distance, a fundamental task in comparative genomics where one seeks a fourth circular genome that minimizes the sum of DCJ distances to three given genomes. While the problem is known to be NP‑complete in general, the authors identify a structural parameter that yields tractability: the number of vertices of degree three in the breakpoint graph constructed from the three input genomes.
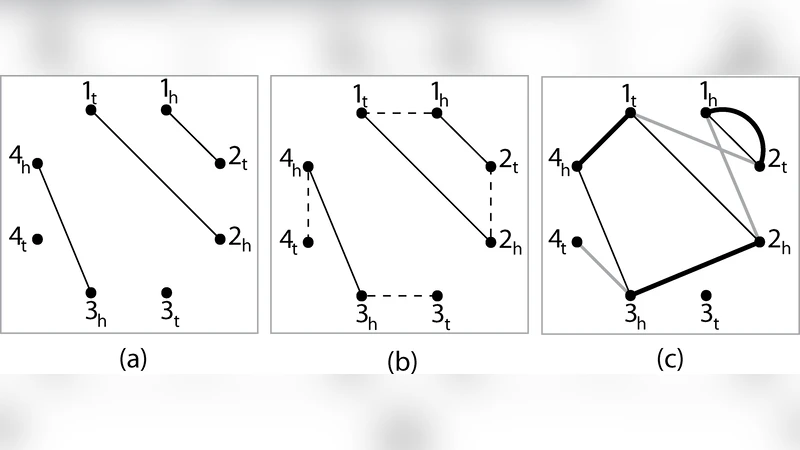
The authors first formalize genomes as matchings on gene extremities and define the breakpoint graph B(G₁,G₂,G₃) as the edge‑colored union of the three genome matchings. In this graph each vertex can be incident to up to three colored edges (one per genome). The DCJ distance between a genome G and a candidate median M equals n minus the number of alternating cycles in B(G,M). Consequently, a circular median maximizes the total number of alternating cycles across the three pairwise breakpoint graphs.
A key technical tool is the “shrinking” operation: given a pair of vertices {u,v}, all edges between them are removed, identical‑colored incident edges are identified, and the two vertices are deleted, producing a smaller graph B·{u,v}. Proposition 1 shows that if a median contains the edge uv, then the number of alternating cycles in the original graph equals the number in the shrunken graph plus the number k of colored edges between u and v. This allows one to contract parts of the graph while preserving optimality information.
The authors first solve the easy case where the breakpoint graph has maximum degree two (i.e., it consists only of even cycles and paths). Lemma 1 proves that for any subgraph H that is a path Pₖ or an even cycle C₂ₖ, the maximum number of alternating cycles is at least |E(H)|/2. Using this bound, a polynomial‑time algorithm can construct an optimal median by independently matching vertices inside each component.
The main contribution concerns vertices of degree three. Let m denote the total number of degree‑3 vertices and ℓ the number of edges whose both endpoints are degree‑3. For each degree‑3 vertex there are at most three ways to choose which two of its three incident colored edges will belong to the median (the third edge must be a median edge). Thus the total number of possible configurations is bounded by 3^m, and each configuration determines a set of edges that will be contracted. After contracting all chosen pairs, the resulting graph contains only degree‑2 components, to which the polynomial algorithm for the easy case applies.
The overall algorithm enumerates all (ℓ+1)·(3·m·m^{2ℓ}+1) configurations, performs O(n³) work per configuration (dominated by matching and cycle counting), and returns the best median found. The running time is therefore O(n³·(ℓ+1)·(3·m·m^{2ℓ}+1)). When m (or ℓ) is treated as a fixed parameter, the algorithm runs in polynomial time, establishing that the DCJ circular median problem is Fixed‑Parameter Tractable (FPT) with respect to the number of degree‑3 vertices.
The paper’s significance lies in providing the first explicit, non‑trivial tractable class for the DCJ median problem. It shows that the computational hardness stems primarily from the presence of degree‑3 vertices in the breakpoint graph. By bounding this structural parameter, the otherwise intractable problem becomes efficiently solvable. The authors also discuss that if m is unbounded, one can remove a limited number of edges incident to degree‑3 vertices to obtain a reduced instance with bounded m, solve it in polynomial time, and then reconstruct a solution for the original instance.
Limitations include the exponential dependence on m and ℓ, which may still be prohibitive for highly rearranged genomes where many degree‑3 vertices appear. The work focuses exclusively on circular genomes; extensions to linear or mixed chromosome models are left for future research. Nonetheless, the theoretical framework and the shrinking technique provide a solid foundation for designing practical heuristics and for further parameterized studies of genome median problems.
Comments & Academic Discussion
Loading comments...
Leave a Comment