Maximum entropy models for antibody diversity
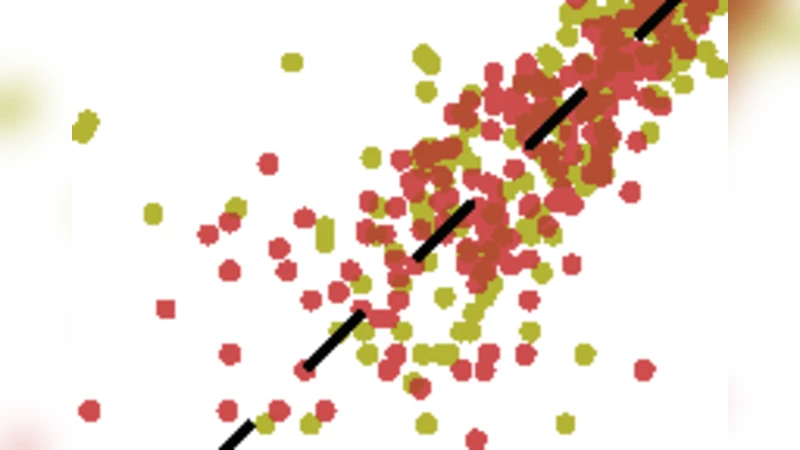
Recognition of pathogens relies on families of proteins showing great diversity. Here we construct maximum entropy models of the sequence repertoire, building on recent experiments that provide a nearly exhaustive sampling of the IgM sequences in zebrafish. These models are based solely on pairwise correlations between residue positions, but correctly capture the higher order statistical properties of the repertoire. Exploiting the interpretation of these models as statistical physics problems, we make several predictions for the collective properties of the sequence ensemble: the distribution of sequences obeys Zipf’s law, the repertoire decomposes into several clusters, and there is a massive restriction of diversity due to the correlations. These predictions are completely inconsistent with models in which amino acid substitutions are made independently at each site, and are in good agreement with the data. Our results suggest that antibody diversity is not limited by the sequences encoded in the genome, and may reflect rapid adaptation to antigenic challenges. This approach should be applicable to the study of the global properties of other protein families.
💡 Research Summary
The paper presents a quantitative framework for describing the immense diversity of antibody repertoires, focusing on IgM sequences from zebrafish. Using an almost exhaustive collection of these sequences, the authors construct a maximum‑entropy (MaxEnt) statistical model that incorporates only two types of empirical constraints: (1) the marginal frequency of each amino‑acid at every position (first‑order statistics) and (2) the pairwise correlations between residues at different positions (second‑order statistics). By maximizing entropy subject to these constraints, they obtain a Boltzmann‑type probability distribution over the space of possible sequences:
P(σ) = Z⁻¹ exp
Comments & Academic Discussion
Loading comments...
Leave a Comment