COLoR - Coordinated On-Line Rankers for Network Reconstruction
Predicting protein interactions is one of the more interesting challenges of the post-genomic era. Many algorithms address this problem as a binary classification problem: given two proteins represented as two vectors of features, predict if they interact or not. Importantly however, computational predictions are only one component of a larger framework for identifying PPI. The most promising candidate pairs can be validated experimentally by testing if they physical bind to each other. Since these experiments are more costly and error prone, the computational predictions serve as a filter, aimed to produce a small number of highly promising candidates. Here we propose to address this problem as a ranking problem: given a network with known interactions, rank all unknown pairs based on the likelihood of their interactions. In this paper we propose a ranking algorithm that trains multiple inter-connected models using a passive aggressive on-line approach. We show good results predicting protein-protein interactions for post synaptic density PPI network. We compare the precision of the ranking algorithm with local classifiers and classic support vector machine. Though the ranking algorithm outperforms the classic SVM classification, its performance is inferior compared to the local supervised method.
💡 Research Summary
The paper addresses the problem of predicting protein‑protein interactions (PPIs) by reframing it from a binary classification task to a ranking problem. Traditional approaches train a single classifier on all protein pairs, but this can over‑generalize because the predictive power of features (e.g., gene expression, domain composition, phylogenetic profiles) varies across different regions of the interaction network. To capture this heterogeneity, the authors propose COLoR (Coordinated On‑line Rankers), a multi‑task learning framework that builds a separate local scoring model for each protein while also maintaining a global model shared by all proteins.
The scoring function combines a local model (W_i) and a global model (W_0) as follows:
(S(p,q)=\beta,p^{\top}W_i q+(1-\beta),p^{\top}W_0 q), where (\beta) balances local versus global contributions. For regularization, the method enforces similarity between the parameters of neighboring proteins in the known interaction graph, using an L2 penalty (\alpha_{ij}|W_i-W_j|^2). The authors set (\alpha_{ij}=1) for edges present in the graph, encouraging adjacent nodes to have similar models. An additional temporal regularizer keeps each model close to its previous value, which stabilizes online updates.
Because the number of proteins (and thus models) can be large, the authors adopt an online Passive‑Aggressive (PA) learning scheme. At each iteration a protein (p_i) is sampled together with a positive partner (p^+) (known interaction) and a negative partner (p^-) (non‑interaction). The hinge loss for the triplet is (l_W(p_i,p^+,p^-)=\max{0,1-S(p_i,p^+)+S(p_i,p^-)}). The PA update solves a constrained quadratic problem that minimizes the sum of the neighbor‑similarity penalties, the deviation from the previous global model, and a slack variable weighted by a constant (C). The closed‑form solution yields an update scalar (\tau = \min{C, l_W/( (1-\beta+N_e\beta)|V|^2 )}) where (V = p_i(p^+ - p^-)^{\top}) and (N_e) is the number of neighbors (including a pseudo‑neighbor representing the previous state). The local model is updated as (W_i \leftarrow \frac{1}{N_e}\sum_{j\in N(i)}W_j + \frac{1}{N_e}\tau V) and the global model as (W_0 \leftarrow W_0 + \tau V). If the loss is zero, no update occurs, making the algorithm computationally efficient.
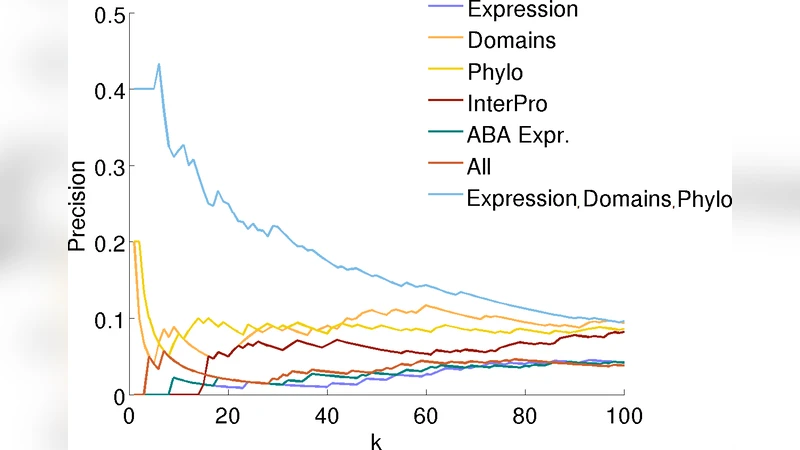
The experimental evaluation uses a curated post‑synaptic density (PSD) PPI network comprising 114 proteins and 211 interactions. Features are assembled from four sources: (1) brain‑related microarray expression (459 GEO samples, normalized and Z‑scored), (2) phylogenetic profiles from Inparanoid (99 species, ortholog scores), (3) protein domains (Pfam‑A) and signatures (InterPro) represented with TF‑IDF weighting, and (4) region‑specific expression from the Allen Brain Atlas (17 brain structures). The authors test several feature combinations and find that the best performance is achieved by merging expression, domain, signature, and phylogenetic data; the Allen Brain Atlas data alone is non‑predictive and degrades performance when combined with the others.
Three predictive strategies are compared: (i) a global linear SVM trained on all pairs, (ii) local SVMs trained independently for each protein (the “local” approach from Bleakley et al., 2007), and (iii) the proposed COLoR algorithm. Performance is measured via 5‑fold cross‑validation, evaluating precision at the top‑k ranked candidate pairs (precision@k). Results show that the global SVM yields the lowest precision across all k. COLoR outperforms the global SVM, especially within the top 40 predictions, but the local SVM consistently achieves the highest precision for every k. This suggests that when sufficient labeled interactions are available for each protein, fully independent models capture the most discriminative patterns; however, COLoR’s ability to share information among neighboring proteins provides a useful middle ground when data are sparse.
The authors discuss several limitations. First, the current experiments involve a relatively small network; scaling to thousands of proteins may increase the computational burden of the neighbor‑similarity regularization. Second, the method relies on hyper‑parameters (\beta) and (C) whose sensitivity is not thoroughly explored, implying that careful tuning would be required in practice. Third, the superiority of local SVMs may be due to the relatively dense labeling in the PSD dataset; in sparser networks, COLoR’s collaborative learning could be more advantageous. Finally, the lack of predictive power from the Allen Brain Atlas highlights that not all biologically relevant data automatically improve PPI prediction; feature relevance must be assessed empirically.
In conclusion, the paper introduces a novel online multi‑task ranking framework for network reconstruction that leverages both local specificity and global consistency. While it does not surpass the best performing local classifiers on the tested dataset, COLoR demonstrates that coordinated online learning can yield better results than a naïve global model and offers a scalable approach for situations where per‑node training data are limited. Future work could focus on extending the method to larger interactomes, automating hyper‑parameter selection, and integrating additional domain‑specific features to further boost predictive accuracy.
Comments & Academic Discussion
Loading comments...
Leave a Comment