Codon Capture and Ambiguous Intermediate Scenarios of Genetic Code Evolution
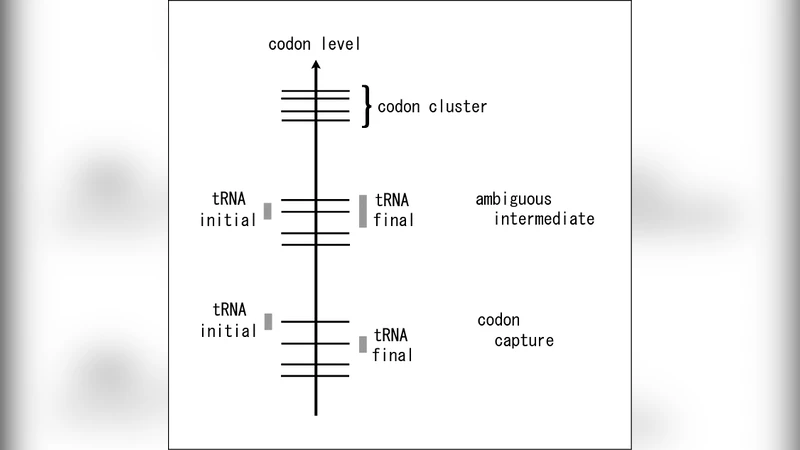
Using the shape space of codons and tRNAs we give a physical description of the genetic code evolution on the basis of the codon capture and ambiguous intermediate scenarios in a consistent manner. In the lowest dimensional version of our description, a physical quantity, codon level is introduced. In terms of the codon levels two scenarios are typically classified into two different routes of the evolutional process. In the case of the ambiguous intermediate scenario we perform an evolutional simulation implemented cost selection of amino acids and confirm a rapid transition of the code change. Such rapidness reduces uncomfortableness of the non-unique translation of the code at intermediate state that is the weakness of the scenario. In the case of the codon capture scenario the survival against mutations under the mutational pressure minimizing GC content in genomes is simulated and it is demonstrated that cells which experience only neutral mutations survive.
💡 Research Summary
**
The paper presents a unified physical framework for two competing hypotheses on the evolution of the genetic code: the codon‑capture scenario and the ambiguous‑intermediate scenario. The authors adopt the concept of a “shape space”—originally used in immunology to describe antigen‑antibody interactions—to model the molecular recognition between codons and transfer RNAs (tRNAs). In this space, each codon is represented by a scalar quantity called the “codon level.” Codon levels cluster into groups that correspond to the family boxes of the standard genetic code, with gaps separating different families. The width of a tRNA’s recognition range along the codon‑level axis reflects its ability to bind one or more codons.
The key distinction between the two evolutionary routes lies in where the mutation occurs in the tRNA molecule. A mutation in the anticodon changes the centre of the recognition range, effectively “jumping” the tRNA from one codon level to another; this is identified with the codon‑capture scenario. Mutations elsewhere (e.g., base modifications, RNA editing) alter only the width of the recognition range while leaving its centre unchanged, allowing a single tRNA to recognize multiple codons simultaneously—this corresponds to the ambiguous‑intermediate scenario.
Selection pressures are incorporated differently for each scenario. In the ambiguous‑intermediate case, the authors assume that natural selection favours lower‑cost amino‑acid synthesis; thus, tRNAs that deliver cheaper amino acids replicate faster. In the codon‑capture case, only cells that experience neutral mutations (i.e., mutations that do not affect the overall GC content) survive under mutational pressure that drives genomes toward lower GC content.
A computational simulation is built around four procedural steps: (1) random alteration of tRNA recognition widths using a weight function; (2) elimination of tRNAs whose widths fall outside biologically plausible bounds; (3) replication of surviving tRNAs with rates proportional to the synthesis cost of their associated amino acids; (4) cell division when the total tRNA population reaches a threshold, after which one daughter cell is followed. The model is deliberately minimal, focusing on two codons (codon‑1 and codon‑2) and two corresponding tRNA species (tRNA‑1 and tRNA‑2). Initially each codon is recognized by its dedicated tRNA. Mutations gradually broaden the recognition range of tRNA‑1 until it can also bind codon‑2. Once this “dual‑recognition” tRNA appears, the takeover of codon‑2 from tRNA‑2 proceeds rapidly, typically within about ten generations for the parameter set used (N≈500). The speed of this transition is largely independent of the exact mutation rate; the mutation rate mainly determines when the takeover begins. The simulation also shows that the variance in recognition‑range width for tRNA‑1 spikes during the transition, then stabilises after the takeover, whereas tRNA‑2’s variance remains low until it disappears.
Although the paper does not present explicit simulations for the codon‑capture scenario, it argues that under a mutational bias toward reduced GC content, only cells that experience neutral mutations (i.e., that do not alter codon usage dramatically) persist, which aligns with empirical observations of stop‑codon reassignment to amino acids in mitochondria and some prokaryotes.
The study’s major contribution is the introduction of the codon‑level scalar as a unifying physical descriptor, allowing both scenarios to be visualized as different trajectories within the same shape space: intra‑cluster transitions for ambiguous‑intermediate evolution and inter‑cluster jumps for codon capture. This provides a conceptual bridge between previously competing models and offers a mechanistic explanation for why ambiguous‑intermediate transitions can be swift enough to avoid prolonged periods of non‑unique translation.
Nevertheless, the model’s simplifications limit its biological realism. Reducing the high‑dimensional properties of codons to a single scalar ignores important aspects such as base‑pairing energetics, wobble rules, and the influence of tRNA‑modifying enzymes. Moreover, the exclusion of other critical components—amino‑acyl‑tRNA synthetases, ribosomal factors, and release factors—means that the model captures only a subset of the forces shaping codon reassignment. Future work could extend the shape space to multiple dimensions, incorporate additional molecular actors, and perform quantitative simulations of the codon‑capture pathway to test the robustness of the conclusions. Despite these caveats, the paper offers a novel perspective that integrates physical modeling with evolutionary genetics, opening avenues for more comprehensive computational studies of genetic‑code evolution.
Comments & Academic Discussion
Loading comments...
Leave a Comment