Log-Normal Distribution of Single Molecule Fluorescence Bursts in Micro/Nano-Fluidic Channels
The width and shape of photon burst histograms pose significant limitations to the identification of single molecules in micro/nano-fluidic channels, and the nature of these histograms is not fully understood. To reach a deeper understanding, we performed computer simulations based on a Gaussian beam intensity profile with various fluidic channel diameters and assuming (i) a deterministic (noise-free) case, (ii) photon emission/absorption noise, and (iii) photon noise with diffusion. Photon noise in narrow channels yields a Gaussian burst distribution while additional strong diffusion produces skewed histograms. We use the fluctuating residence time picture [Phys. Rev. Lett. 80, 2386-2388 (1998)] and conclude that the skewness of the photon number distribution is caused by the longitudinal diffusive component of the motion of the molecules as they traverse the laser beam. In the case of strong diffusion in narrow channels, this effect leads to a log-normal distribution. We show that the same effect can transform the separate peaks of the photon burst histograms of multiple molecule mixtures into a single log-normal shape.
💡 Research Summary
This paper investigates the statistical shape of photon‑burst histograms obtained in single‑molecule fluorescence experiments performed in micro‑ and nano‑fluidic channels. The authors focus on the MAPS (Microfluidic System for Analyzing Proteins in a Single Complex) platform, where quantum‑dot‑labeled proteins drift through a Gaussian laser beam, emit a burst of photons, and the burst size is used to identify the molecule. In practice, the burst‑size histograms are broadened and often skewed, limiting the ability to discriminate different species.
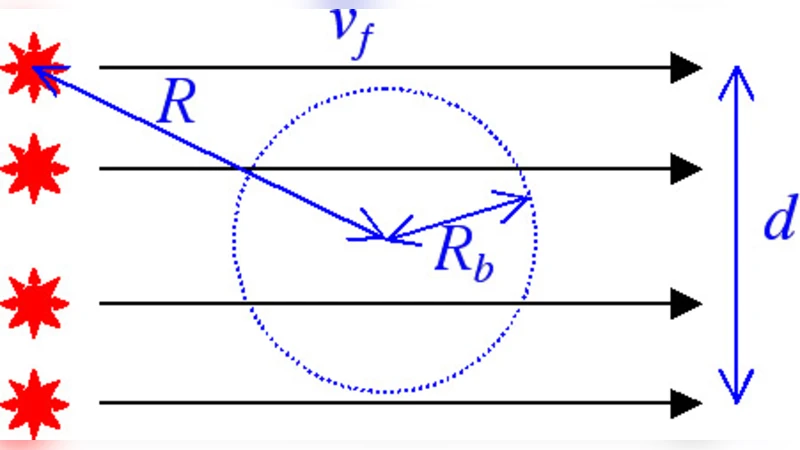
To elucidate the origin of this broadening, the authors conduct Monte‑Carlo simulations that model three distinct situations: (i) a deterministic, noise‑free case (no photon shot noise, no diffusion), (ii) photon shot noise only (Poisson statistics for excitation and emission), and (iii) photon shot noise combined with molecular diffusion. The laser intensity follows a Gaussian profile with effective radius (R_b = 320) arbitrary units. Molecules move with a constant drift velocity (laminar flow) superimposed with a random walk characterized by a diffusion strength parameter (D), defined as the squared ratio of the diffusive displacement to the beam diameter over the time required to cross the beam.
In the deterministic case, the histogram collapses to a single line, confirming that without any stochastic element the burst size would be perfectly reproducible. When only photon shot noise is present, the histogram becomes Gaussian. This agrees with the central‑limit theorem: the sum of many independent Poisson events (photon absorptions and emissions) yields a normal distribution, and the simulated normal probability plot is a straight line over the 1–99 % range.
When diffusion is introduced, the situation changes dramatically. Diffusion causes fluctuations in the residence time of a molecule inside the excitation zone. Longer residence times expose the molecule to higher laser intensity for a longer period, producing a larger photon count; shorter residence times produce fewer photons. Because the residence time appears multiplicatively in the total photon number, the logarithm of the photon count tends toward a Gaussian distribution, i.e., the photon‑burst size follows a log‑normal law.
The authors explore two channel geometries: a narrow channel where the beam radius is sixteen times the channel half‑width ((R_b/d = 16)) and a wide channel where the ratio is 0.32. In the narrow channel with strong diffusion ((D = 5.63)), the burst‑size histogram is strongly right‑skewed and the log‑normal probability plot is linear over the 1–99 % range, indicating a clear log‑normal behavior. In the wide channel with the same diffusion strength, the histogram is broader and the log‑normality holds only over a reduced range (10–99 %), showing that channel width degrades the ideal log‑normal shape by adding geometric variability to the residence time.
The paper also examines mixtures of three molecular species, each producing a distinct mean photon count (≈100, 200, 300 photons). Without diffusion the histogram displays three well‑separated peaks. Even weak diffusion ((D = 0.13)) begins to merge the second and third peaks, and strong diffusion ((D = 1.6)) collapses all three peaks into a single, approximately log‑normal distribution. This demonstrates that diffusion can erase the discrete signatures of different labels, a critical limitation for multiplexed single‑molecule assays.
Overall, the study links the observed skewness and width of photon‑burst histograms to residence‑time fluctuations caused by diffusion, extending the “fluctuating residence time” concept originally introduced for nanoparticle growth in gas evaporation (Söderlund et al., PRL 80, 2386, 1998). The findings provide practical guidance: to preserve Gaussian or well‑separated peaks, one should minimize diffusion (e.g., by reducing temperature, increasing flow speed, or using narrower channels). Conversely, when diffusion cannot be avoided, the resulting log‑normal distribution can be anticipated and accounted for in data analysis, for instance by fitting log‑normal models or by employing statistical deconvolution techniques. The work thus clarifies a long‑standing discrepancy between ideal Poisson‑based models and experimental burst‑size histograms, and it offers a quantitative framework for designing more reliable single‑molecule fluorescence detection systems in micro‑ and nano‑fluidic environments.
Comments & Academic Discussion
Loading comments...
Leave a Comment