Group Lasso with Overlaps: the Latent Group Lasso approach
We study a norm for structured sparsity which leads to sparse linear predictors whose supports are unions of prede ned overlapping groups of variables. We call the obtained formulation latent group Lasso, since it is based on applying the usual group Lasso penalty on a set of latent variables. A detailed analysis of the norm and its properties is presented and we characterize conditions under which the set of groups associated with latent variables are correctly identi ed. We motivate and discuss the delicate choice of weights associated to each group, and illustrate this approach on simulated data and on the problem of breast cancer prognosis from gene expression data.
💡 Research Summary
**
The paper introduces a novel regularization framework called the Latent Group Lasso, designed to handle overlapping groups of variables in high‑dimensional learning problems. Traditional Lasso (ℓ₁) promotes sparsity at the individual variable level, while the classical Group Lasso (ℓ₁/ℓ₂) enforces sparsity at the group level but assumes that groups form a partition of the feature set. When groups overlap, the standard Group Lasso no longer guarantees that the selected support is a union of whole groups; instead, it often yields supports that are intersections of complements of groups, which is undesirable for many structured‑sparsity applications.
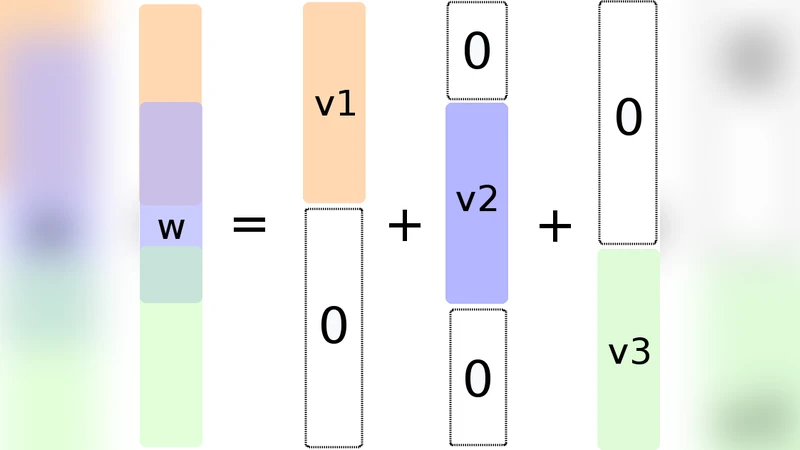
To overcome this limitation, the authors propose to introduce, for each predefined group g ∈ G, a latent vector v⁽ᵍ⁾ whose support is constrained to lie inside g. The original parameter vector w is expressed as the sum of all latent vectors, w = ∑₍g∈G₎ v⁽ᵍ⁾. The regularizer is then defined as the ℓ₁ norm of the ℓ₂ norms of these latent vectors, weighted by positive scalars d_g:
Ω_{G∪}(w) = min_{v∈V_G, ∑v⁽ᵍ⁾=w} ∑_{g∈G} d_g ‖v⁽ᵍ⁾‖₂.
Applying the ℓ₁/ℓ₂ penalty to the latent variables forces entire groups to be either completely active (v⁽ᵍ⁾ ≠ 0, implying all variables in g can be non‑zero) or completely inactive (v⁽ᵍ⁾ = 0, forcing all variables in g to be zero). Consequently, the support of the estimated ŵ is guaranteed to be a union of selected groups, matching the desired structured sparsity pattern.
The paper provides a thorough mathematical analysis of Ω_{G∪}. It proves that Ω_{G∪} is indeed a norm, derives several equivalent variational formulations, and shows that regularizing with Ω_{G∪} is equivalent to a covariate duplication scheme where each group has its own copy of the variables and a standard Group Lasso is applied to the duplicated set. This equivalence enables the use of existing proximal‑gradient, ADMM, or block‑coordinate descent algorithms without major modifications. Moreover, the authors connect Ω_{G∪} to multiple kernel learning, interpreting the latent groups as separate kernels whose ℓ₁‑weighted combination is penalized.
A central theoretical contribution is the notion of group‑support: the set of groups whose latent vectors are non‑zero. This is a stronger concept than ordinary support recovery because it directly reflects the underlying group structure. Using tools from high‑dimensional statistics and consistency theory, the authors establish sufficient conditions under which the estimator obtained by minimizing a convex loss plus λ Ω_{G∪} consistently recovers the true group‑support as the sample size grows. The conditions involve incoherence-type assumptions on the design matrix and appropriate scaling of the regularization parameter λ.
Weight selection (the d_g’s) is highlighted as a critical practical issue. In the overlapping setting, naive uniform weights can bias the solution toward larger groups or cause excessive shrinkage of small groups. The authors recommend scaling weights by the inverse square root of group size (d_g = 1/√|g|) as a default, and discuss alternative data‑driven schemes that adapt to the degree of overlap and correlation structure.
Empirical validation is performed on two fronts. First, synthetic experiments illustrate that the Latent Group Lasso achieves higher support‑recovery rates, lower estimation error, and more stable performance across different noise levels compared with the standard Lasso, the original overlapping Group Lasso (Jenatton et al., 2009), and other structured sparsity methods. The experiments also demonstrate the sensitivity to weight choices, confirming the theoretical guidance. Second, the method is applied to a breast‑cancer prognosis problem using gene‑expression data. Groups are defined from biological pathways and protein‑protein interaction networks. The Latent Group Lasso selects entire pathways rather than isolated genes, leading to models that are more interpretable and that achieve better predictive accuracy in cross‑validation than models built with unstructured Lasso or with standard Group Lasso on a partition of genes.
In summary, the paper makes the following contributions: (1) a new norm Ω_{G∪} that enforces union‑of‑groups sparsity via latent variables; (2) a rigorous analysis of its mathematical properties, equivalence to covariate duplication, and connection to multiple kernel learning; (3) the introduction of group‑support and consistency results for its recovery; (4) an extensive discussion of weight design for overlapping groups; and (5) thorough experimental evidence on both synthetic and real biomedical data. The work opens avenues for further extensions to non‑linear models, multi‑task learning, and more complex graph‑based structures.
Comments & Academic Discussion
Loading comments...
Leave a Comment