Toward tunable RNA thermo-switches for temperature dependent gene expression
RNA thermometers are mRNA strands with a temperature dependent secondary structure: depending on the spatial conformation, the mRNA strand can get translated (on-state) or can be inaccessible for ribosomes binding (off-state). These have been found in a number of microorganisms (mainly pathogens), where they are used to adaptively regulate the gene expression, in response to changes in the environmental temperature. Besides naturally occurring RNA thermometers, synthetic RNA thermometers have been recently designed by modifying their natural counterparts (Hofacker et al., 2003). The newly designed RNA thermometers are simpler, and exhibit a sharper switching between off- and on-states. However, the proposed trial-and-error design procedure has not been algorithmically formalized, and the switching temperature is rigidly determined by the natural RNA thermometer used as template for the design. We developed a general algorithmic procedure (consensus distribution) for the design of RNA thermo-switches with a tunable switching temperature that can be decided in advance by the designer. A software tool with a user friendly GUI has been written to automate the design of RNA thermo-switches with a desired threshold temperature. Starting from a natural template, a new RNA thermometer has been designed by our method for a new desired threshold temperature of 32C. The designed RNA thermo-switch has been experimentally validated by using it to control the expression of lucifarase. A 9.2 fold increase of luminescence has been observed between 30C and 37C, whereas between 20C and 30C the luminescence increase is less than 3-fold. This work represents a first step towards the design of flexible and tunable RNA thermometers that can be used for a precise control of gene expression without the need of external chemicals and possibly for temperature measurements at a nano-scale resolution.
💡 Research Summary
RNA thermometers are messenger RNAs whose secondary structure changes with temperature, thereby exposing or occluding the ribosome binding site (RBS) and controlling translation. Naturally occurring thermometers, especially the ROSE family, function in a narrow temperature window around 37 °C and have been co‑opted in synthetic biology to achieve temperature‑dependent gene expression without chemical inducers. However, existing design approaches rely on trial‑and‑error modifications of natural sequences, offering little control over the desired switching temperature.
In this work the authors first performed a systematic analysis of 32 known ROSE RNA thermometers using RNAfold at 37 °C. They calculated the cumulative Gibbs free‑energy (ΔG) along each sequence and discovered a highly conserved profile: a high‑energy plateau covering the hairpin that contains the RBS, followed by a linear decline downstream. This “consensus Gibbs energy distribution” reflects the balance between sufficient instability (to allow opening at higher temperature) and enough stability (to keep the RBS hidden at lower temperature).
The central design principle is to reproduce this consensus profile at a user‑specified temperature (T_target). Because ΔG is temperature‑dependent, the nucleotide sequence must be altered so that, at T_target, the free‑energy landscape matches the 37 °C consensus. The authors formalized this as an algorithm that builds the hairpin loop in 4‑nt blocks. For each block the algorithm enumerates all 36 possible loop sequences, filters them according to (i) compatibility with the two terminal bases of the previously placed block, (ii) compliance with fixed nucleotides required by the scar, RBS, start codon, and (iii) the overall ΔG tolerance set by the user. The remaining candidates are scored by how closely their cumulative ΔG follows the consensus curve; the best‑scoring block is appended and the process repeats until the full hairpin is constructed.
The algorithm was implemented in a user‑friendly graphical tool (RNAthermoDesigner). Users input the desired switching temperature and an allowable ΔG deviation (e.g., ±2 kcal/mol). The software then outputs all feasible sequences. When the tolerance was set to 2 kcal/mol for a target of 32 °C, 69 viable sequences were generated; with an unrestricted tolerance, 1 536 sequences (the full combinatorial space for the given length) were produced, most of which were too stable or too unstable to act as effective thermometers. The current version only handles stacked loops; inclusion of bulges and internal loops is planned for future releases, which should increase design flexibility.
To validate the method, the authors selected a natural ROSE thermometer as a template and used the algorithm to design a new hairpin with a predicted switching temperature of 32 °C. The resulting construct (registered as BBa_K115017 in the iGEM parts registry) was synthesized, cloned upstream of a Renilla luciferase gene, and transformed into Escherichia coli. Luminescence was measured at 20 °C, 30 °C, and 37 °C. Between 30 °C and 37 °C the engineered strain showed a 9.2‑fold increase in luciferase activity, whereas the increase between 20 °C and 30 °C was only 1.9‑fold, confirming that the hairpin remains largely repressive below ~32 °C and opens sharply above it. A control construct with a temperature‑insensitive RBS displayed only a modest 2.5‑fold increase across the same temperature range, underscoring the specific effect of the designed thermometer.
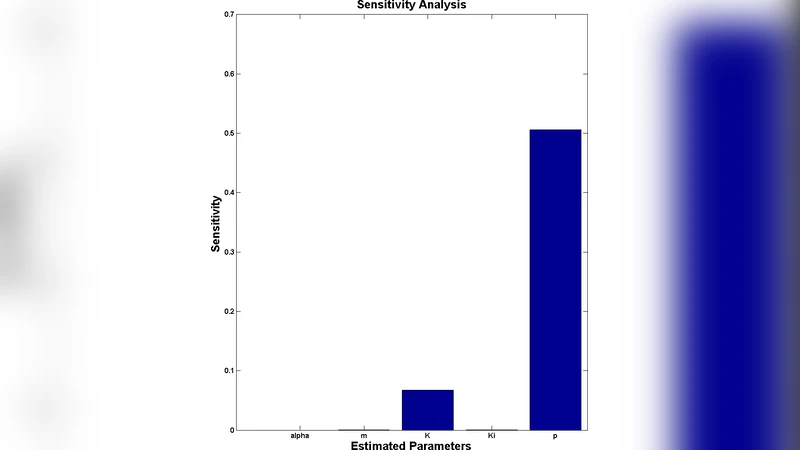
Beyond experimental validation, the authors constructed a mathematical model of temperature‑dependent enzyme production. The model assumes constitutive transcription of the luciferase mRNA, a Hill‑type activation function describing temperature‑driven translation efficiency, and an additional term for thermal inactivation of the enzyme at high temperatures. Parameters (K, m, p, etc.) were fitted to the measured data using a genetic algorithm. The fitted K values (≈28.8 for the 30 °C design and ≈33.1 for the 37 °C design) correspond closely to the intended switching temperatures, confirming that the model captures the essential dynamics. Sensitivity analysis identified the parameter p, governing the “off‑switch” hardness at elevated temperatures, as the most influential factor, suggesting that fine‑tuning of high‑temperature repression could further sharpen the response.
Overall, the study delivers three major contributions: (1) a quantitative, free‑energy‑based specification for RNA thermometer design, (2) an automated algorithm and software that translate a desired temperature into concrete nucleotide sequences while respecting biological constraints, and (3) experimental and computational validation that the designed thermometers function as intended. Limitations include the current restriction to stacked‑loop architectures and the focus on the ROSE family; extending the consensus analysis to other families (PrfA, FourU) and incorporating more diverse secondary‑structure motifs will broaden applicability. Moreover, the work was performed in a single bacterial host; variations in intracellular ion concentrations, RNA‑binding proteins, and metabolic state could shift the effective switching temperature in other organisms, a factor that warrants further investigation.
In conclusion, this paper establishes a systematic, reproducible pipeline for engineering tunable RNA thermometers, moving the field beyond ad‑hoc trial‑and‑error methods toward rational, predictive design. The approach opens the door to temperature‑controlled gene circuits, biosensors with nanometer‑scale spatial resolution, and synthetic biology applications where chemical inducers are undesirable or impractical. Future work expanding structural diversity and testing across multiple chassis will solidify RNA thermometers as a versatile tool in the synthetic biologist’s repertoire.
Comments & Academic Discussion
Loading comments...
Leave a Comment