Bias Plus Variance Decomposition for Survival Analysis Problems
Bias - variance decomposition of the expected error defined for regression and classification problems is an important tool to study and compare different algorithms, to find the best areas for their application. Here the decomposition is introduced for the survival analysis problem. In our experiments, we study bias -variance parts of the expected error for two algorithms: original Cox proportional hazard regression and CoxPath, path algorithm for L1-regularized Cox regression, on the series of increased training sets. The experiments demonstrate that, contrary expectations, CoxPath does not necessarily have an advantage over Cox regression.
💡 Research Summary
The paper introduces a bias‑variance decomposition framework for survival analysis, extending a concept traditionally applied only to regression and classification problems. Survival data consist of a covariate vector x, a survival time t, and a censoring indicator δ. Model performance is usually measured by Harrell’s concordance index (C‑index), which evaluates how well predicted risk scores order the observed failure times. By interpreting the C‑index as the proportion of correctly ordered pairs (assuming continuous features and negligible ties), the authors map the survival problem onto a binary classification setting, allowing them to reuse the classic bias‑variance decomposition for zero‑one loss:
E(C) = 0.5·bias²(x) + 0.5·variance²(x) + 0.5·σ²(x).
Here bias² quantifies the systematic deviation between the distribution of predictions obtained from different training sets of a fixed size and the true label distribution; variance² measures the variability of predictions across those training sets; σ² captures irreducible noise inherent in the data.
Two Cox‑type models are compared: (1) the standard Cox proportional hazards (PH) regression and (2) CoxPath, an L1‑regularized path algorithm that fits a sequence of models for different regularization strengths λ and selects the best one according to a performance criterion. Regularization is expected to reduce variance (making the model more robust to small‑sample fluctuations) while model selection may increase bias, leading to a classic bias‑variance trade‑off.
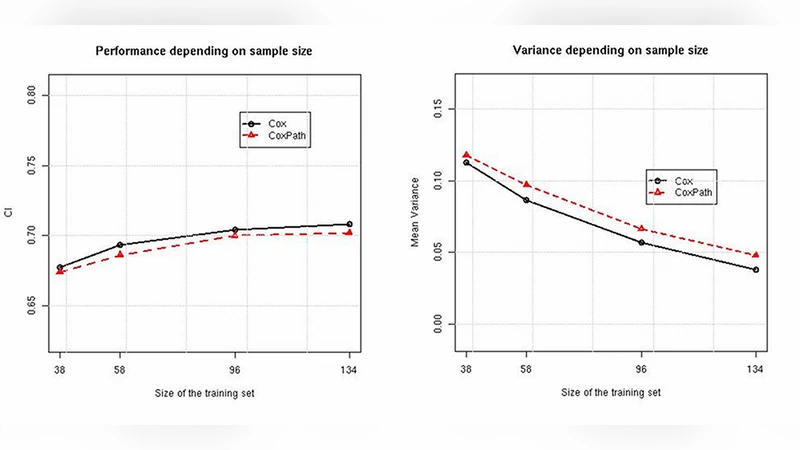
Empirical evaluation uses two real‑world datasets. The PBC dataset (17 features, 228 patients) originates from a primary biliary cirrhosis trial, while the Ro02s dataset (7 aggregated features derived from 7,399 gene expressions) concerns lymphoma patients. For each dataset, 80 % of the observations are held out as a test set. From the remaining 20 %, training subsets of increasing size are drawn randomly; for each size, 20 distinct training sets are generated, and the whole procedure is repeated ten times with different test splits. Models trained on each training set are evaluated on the single test set, yielding estimates of variance and overall error (1 – E(C)).
Results show that bias is virtually identical for both algorithms across all training sizes, confirming that the primary differences stem from variance and total error. On the PBC data, CoxPath achieves lower variance and better overall performance for every sample size, indicating that regularization successfully stabilizes predictions without incurring a substantial bias penalty. Conversely, on the Ro02s data, CoxPath exhibits higher variance and worse performance than the unregularized Cox PH model, suggesting that the model‑selection step introduces sensitivity to the particular training sample that outweighs the variance‑reduction benefit of regularization.
These findings highlight that L1 regularization does not universally improve survival models; its impact depends on data characteristics such as dimensionality, sample size, and feature structure. The authors propose further experiments with synthetic data and with fixed‑λ L1‑regularized Cox models to disentangle the effects of regularization strength from model‑selection bias. Such investigations could yield practical guidelines for choosing between plain Cox PH regression and regularized variants based on the specific properties of a given survival dataset.
In summary, the paper demonstrates that bias‑variance decomposition can be meaningfully defined for survival analysis, provides a concrete methodology for its computation, and uses it to reveal nuanced trade‑offs between Cox PH and CoxPath. The work offers a valuable diagnostic tool for researchers and practitioners aiming to understand and optimize the error behavior of survival‑prediction algorithms.
Comments & Academic Discussion
Loading comments...
Leave a Comment