Unsupervised K-Nearest Neighbor Regression
In many scientific disciplines structures in high-dimensional data have to be found, e.g., in stellar spectra, in genome data, or in face recognition tasks. In this work we present a novel approach to non-linear dimensionality reduction. It is based on fitting K-nearest neighbor regression to the unsupervised regression framework for learning of low-dimensional manifolds. Similar to related approaches that are mostly based on kernel methods, unsupervised K-nearest neighbor (UNN) regression optimizes latent variables w.r.t. the data space reconstruction error employing the K-nearest neighbor heuristic. The problem of optimizing latent neighborhoods is difficult to solve, but the UNN formulation allows the design of efficient strategies that iteratively embed latent points to fixed neighborhood topologies. UNN is well appropriate for sorting of high-dimensional data. The iterative variants are analyzed experimentally.
💡 Research Summary
The paper introduces a novel unsupervised dimensionality‑reduction technique called Unsupervised K‑Nearest Neighbor (UNN) regression. Building on the general unsupervised regression framework proposed by Meinicke, the authors replace the usual kernel‑based regression function with a simple K‑nearest‑neighbor (KNN) estimator. The goal is to find a low‑dimensional latent matrix X such that, when the KNN regression function is applied to X, the high‑dimensional data matrix Y is reconstructed with minimal data‑space reconstruction error (DSRE):
E(X) = (1/N)‖Y – f_UNN(X)‖F², f_UNN(x;X) = (1/K)∑{i∈N_K(x,X)} y_i.
Because KNN depends only on neighborhood relations, the absolute positions of latent points are irrelevant; only the adjacency structure matters. The authors therefore fix a simple one‑dimensional latent topology (equidistant points on a line) and search for a permutation of the N data points that yields the smallest DSRE. Two greedy insertion heuristics are proposed:
-
UNN‑1 (global insertion) – For each new data point, all N̂ + 1 possible insertion slots between already placed points are evaluated. The slot with the lowest DSRE is chosen, the point is inserted, and the process repeats until all points are placed. The cost of evaluating one slot is O(K·d) (K neighbours, d‑dimensional outputs). Since N̂ + 1 slots are examined, the overall complexity per iteration is O(N·K·d), which simplifies to O(N) because K and d are constants in practice.
-
UNN‑2 (local insertion) – Instead of testing every slot, the algorithm first finds the already‑embedded point y* that is nearest to the new point y in the original data space (cost O(N·d)). Then only the two positions immediately left and right of y* are evaluated. Consequently, each iteration costs O(N·d) for the nearest‑neighbor search plus O(K·d) for the two DSRE evaluations, again yielding an overall O(N) runtime but with a substantially smaller constant factor than UNN‑1.
The authors evaluate both strategies on three benchmark problems:
-
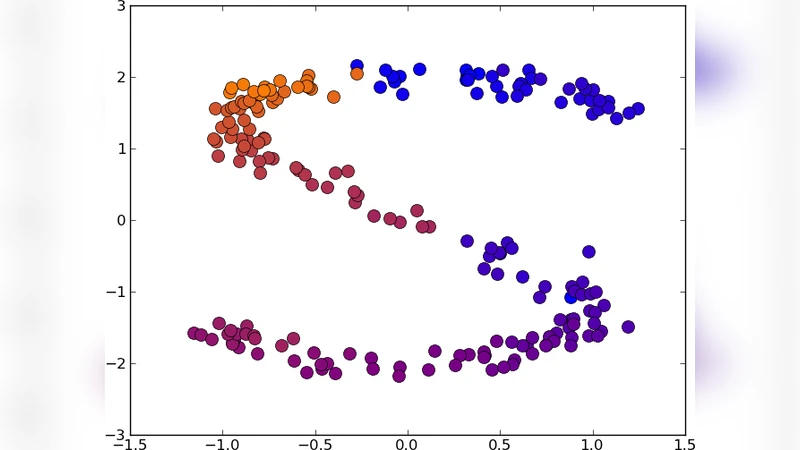
2‑D “S” curve (N = 200) – With K = 5, UNN‑1 achieves a DSRE of 19.6 (initial DSRE ≈ 202) while UNN‑2 reaches 29.2. Both methods correctly order points along the curve, though occasional local optima appear.
-
3‑D “S” curve (N = 500) and a variant with a central hole (N = 400) – For K = 10, UNN‑1 reduces DSRE to 101.9 (hole case 139.0) compared with the initial values of 691.3 and 196.6, respectively. UNN‑2 yields slightly higher DSREs (140.4 and 145.3) but remains far better than the baseline and comparable to Locally Linear Embedding (LLE), which in many cases produces larger errors.
-
USPS handwritten digits (2’s and 5’s, 100 samples each, 256‑dimensional) – Embedding into a one‑dimensional manifold with K = 10, UNN‑2 produces a DSRE of 70.1 for the 2’s (initial ≈ 577) and 145.4 for the 5’s (initial ≈ 197). Visual inspection shows that similar digits cluster together along the latent line, confirming that the method preserves perceptual similarity.
A comprehensive DSRE table (including LLE for comparison) demonstrates that UNN‑1 consistently attains the lowest reconstruction error across all datasets and neighborhood sizes, while UNN‑2 offers a favorable trade‑off between speed and accuracy. The experiments also reveal the expected increase in DSRE as K grows, reflecting the smoothing effect of larger neighborhoods.
The paper concludes that fitting a fast, non‑parametric regression model into the unsupervised setting yields an efficient dimensionality‑reduction tool. Both UNN strategies run in linear time with respect to the number of samples, making them suitable for large‑scale or high‑dimensional problems where kernel‑based methods become prohibitive. However, the current formulation is limited to one‑dimensional latent grids and relies on greedy insertion, which may get trapped in local minima. Future work is outlined to extend the approach to higher‑dimensional latent topologies (e.g., 2‑D grids), to develop stochastic or global optimization schemes (e.g., simulated annealing, genetic algorithms) for better exploration of the permutation space, and to investigate regularization techniques that prevent pathological spreading of latent points.
Comments & Academic Discussion
Loading comments...
Leave a Comment