Tunable kinetic proofreading in a model with molecular frustration
In complex systems, feedback loops can build intricate emergent phenomena, so that a description of the whole system cannot be easily derived from the properties of the individual parts. Here we propose that inter-molecular frustration mechanisms can provide non trivial feedback loops which can develop nontrivial specificity amplification. We show that this mechanism can be seen as a more general form of a kinetic proofreading mechanism, with an interesting new property, namely the ability to tune the specificity amplification by changing the reactants concentrations. This contrasts with the classical kinetic proofreading mechanism in which specificity is a function of only the reaction rate constants involved in a chemical pathway. These results are also interesting because they show that a wide class of frustration models exists that share the same underlining kinetic proofreading mechanisms, with even richer properties. These models can find applications in different areas such as evolutionary biology, immunology and biochemistry.
💡 Research Summary
The paper “Tunable kinetic proofreading in a model with molecular frustration” investigates how inter‑molecular frustration can be understood as a generalized form of kinetic proofreading (KP) and, crucially, how it endows the system with a tunable specificity that depends on reactant concentrations rather than solely on kinetic rate constants. The authors begin by reviewing the classic KP concept introduced by Hopfield (1974) and later applied to T‑cell activation by McKeithan (1995). In the traditional KP framework, a ligand‑receptor pair proceeds through a series of N intermediate steps before reaching a final product. If the dissociation rates of the two competing pathways differ (k‑1 versus k‑1*), the ratio of final products can be amplified exponentially with N, yielding high specificity. However, this amplification is fixed once the kinetic constants are set; specificity cannot be altered without changing the chemistry of the system, and increasing specificity inevitably reduces overall sensitivity.
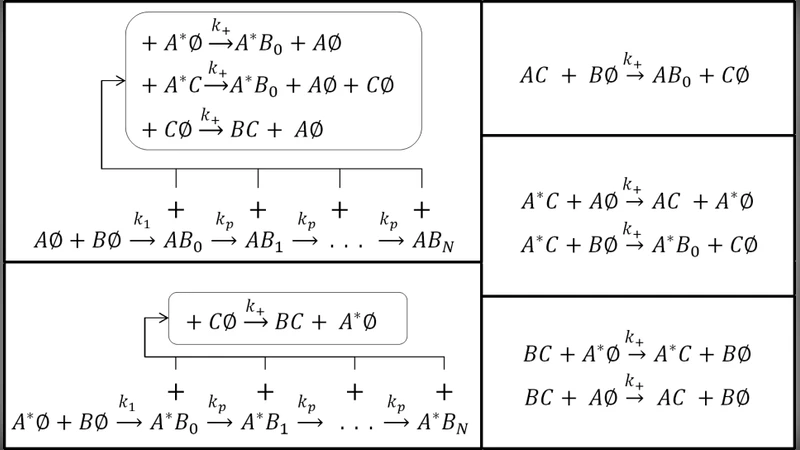
The authors then introduce a “cellular frustration” concept previously used to model assortative mating and speciation, where the presence of other agents destabilizes existing pairings. Translating this idea to molecular interactions, they propose a model with four species: A, A*, B, and C. The interaction hierarchy is encoded in a table (Figure 2) such that A binds B most strongly, B binds C more strongly than A or A*, and A* binds B with intermediate affinity. Crucially, a complex can be destabilized when a third molecule of higher affinity binds to one of its components, mimicking a frustration event. For example, an AB complex can be broken by a free C or by an A* molecule, whereas an A*B complex is only vulnerable to free C. This asymmetry creates a concentration‑dependent effective dissociation rate: the apparent k‑1 for AB becomes k‑1·
Comments & Academic Discussion
Loading comments...
Leave a Comment