Protein multi-scale organization through graph partitioning and robustness analysis: Application to the myosin-myosin light chain interaction
Despite the recognized importance of the multi-scale spatio-temporal organization of proteins, most computational tools can only access a limited spectrum of time and spatial scales, thereby ignoring the effects on protein behavior of the intricate coupling between the different scales. Starting from a physico-chemical atomistic network of interactions that encodes the structure of the protein, we introduce a methodology based on multi-scale graph partitioning that can uncover partitions and levels of organization of proteins that span the whole range of scales, revealing biological features occurring at different levels of organization and tracking their effect across scales. Additionally, we introduce a measure of robustness to quantify the relevance of the partitions through the generation of biochemically-motivated surrogate random graph models. We apply the method to four distinct conformations of myosin tail interacting protein, a protein from the molecular motor of the malaria parasite, and study properties that have been experimentally addressed such as the closing mechanism, the presence of conserved clusters, and the identification through computational mutational analysis of key residues for binding.
💡 Research Summary
The paper presents a novel computational framework for uncovering the hierarchical, multi‑scale organization of proteins by representing them as atomistic interaction networks and applying a Markov‑process‑based community detection method called “Stability.” Starting from a high‑resolution PDB structure, each atom becomes a node and each covalent bond, hydrogen bond, hydrophobic contact, or salt bridge is encoded as a weighted edge whose weight reflects the underlying potential energy of the interaction. This weighted graph therefore retains both chemical specificity and spatial information.
Stability measures how likely a random walk on the graph remains trapped within a set of nodes over a given Markov time t. Small t highlights tightly coupled groups of atoms (e.g., chemical groups or side‑chain contacts), while larger t reveals increasingly coarse communities that correspond to larger structural motifs such as secondary‑structure elements, helix bundles, or whole domains. By sweeping t continuously and optimizing Stability at each point with the fast Louvain heuristic, the authors obtain a hierarchy of partitions that spans the entire range from atomic to domain scale.
A key methodological advance is the introduction of a robustness assessment. The authors generate ensembles of surrogate random graphs that preserve specific biochemical constraints (e.g., degree distribution, interaction type frequencies) and compute the expected Stability for each surrogate. A partition is deemed robust when its observed Stability significantly exceeds the surrogate expectation, providing a statistically grounded way to decide which scales are biologically meaningful.
The framework is further extended with a computational mutagenesis protocol. By virtually removing or weakening a node (or its incident edges) and re‑evaluating the Stability hierarchy, the impact of each residue on the global multi‑scale organization can be quantified. Residues whose perturbation causes a large drop in Stability at any scale are identified as “hot‑spots,” i.e., potential functional or drug‑target sites.
Method validation is performed on Escherichia coli adenylate kinase (AdK), a protein whose opening/closing mechanism is well characterized experimentally. The multi‑scale partitions correctly recover known functional units: short‑time clusters correspond to the phosphate‑binding loop, intermediate‑time clusters to the lid domain, and long‑time clusters to the whole enzyme, mirroring the experimentally observed conformational transition.
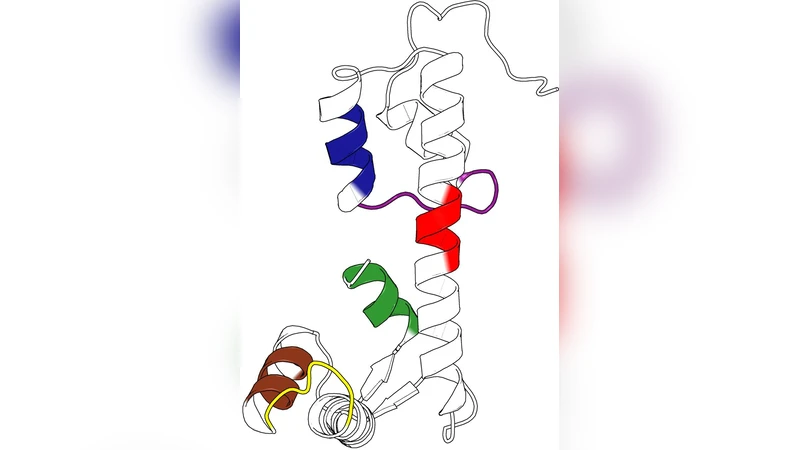
The main biological application focuses on the myosin‑tail interacting protein (MTIP) from Plasmodium species, a key component of the malaria parasite’s motor complex. Four crystal structures are analyzed: two species (P. knowlesi and P. falciparum) each in apo form and bound to a MyoA tail peptide. The analysis reveals:
- At short Markov times, the graph decomposes into chemically coherent groups (hydrogen‑bond networks, side‑chain contacts).
- At intermediate times, the N‑terminal helix and the C‑terminal helix each form distinct communities, reflecting their semi‑independent motions.
- At long times, the two helices merge into a single community, representing the “closing” of MTIP around the MyoA tail. This provides a mechanistic picture of how MTIP wraps the tail.
Robustness testing shows that the C‑terminal community is highly significant compared with surrogate graphs, indicating evolutionary conservation of this functional module. Computational mutagenesis of the MyoA peptide identifies several residues (e.g., Leu‑15, Phe‑12, Ile‑9) whose removal dramatically reduces the Stability of the MTIP‑MyoA complex, pinpointing them as critical binding hot‑spots. These predictions align with experimental binding assays that reported the same residues as essential for high‑affinity interaction.
Overall, the study demonstrates that atomistic graph representations combined with multi‑scale Stability analysis can automatically reveal the full hierarchy of structural organization in proteins, assess the statistical significance of each level, and predict functionally important residues. The approach bridges the gap between fine‑grained molecular detail and coarse‑grained functional insight, offering a powerful tool for structural biology, allostery studies, and rational drug design.
Comments & Academic Discussion
Loading comments...
Leave a Comment