Modeling the early steps of cytoplasmic trafficking in viral infection and gene delivery
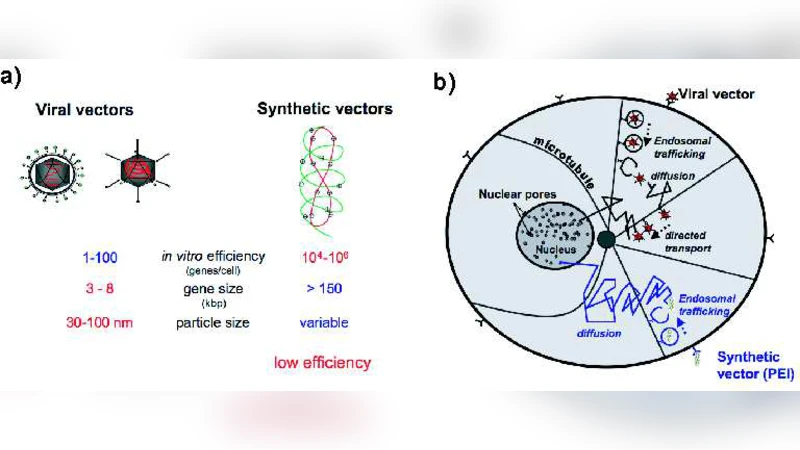
Gene delivery of nucleic acid to the cell nucleus is a fundamental step in gene therapy. In this review of modeling drug and gene delivery, we focus on the particular stage of plasmid DNA or virus cytoplasmic trafficking. A challenging problem is to quantify the success of this limiting stage. We present some models and simulations of plasmid trafficking and of the limiting phase of DNA-polycation escape from an endosome and discuss virus cytoplasmic trafficking. The models can be used to assess the success of viral escape from endosomes, to quantify the early step of viral-cell infection, and to propose new simulation tools for designing new hybrid-viruses as synthetic vectors.
💡 Research Summary
This review article provides a comprehensive quantitative analysis of the early intracellular steps that limit the efficiency of gene delivery, focusing on plasmid DNA and viral particle trafficking within the cytoplasm. The authors first discuss the challenges associated with endosomal escape, a critical barrier for both viral vectors and synthetic carriers such as cationic polymers (e.g., polyethylenimine, PEI). They formalize the widely cited “proton‑spunge” hypothesis using a set of coupled kinetic and osmotic equations: acidification of the endosome leads to protonation of the polymer, increasing its net positive charge, which in turn drives an influx of chloride ions and water to maintain electroneutrality. The resulting osmotic pressure causes swelling and eventual rupture of the endosomal membrane. By assigning realistic values to the protonation rate constants, polymer charge capacity, and membrane elasticity, the authors perform stochastic particle‑based simulations that reproduce experimentally observed pH thresholds and rupture times. Sensitivity analyses reveal that polymer buffering capacity and endosomal membrane rigidity are the dominant determinants of escape efficiency.
The second major component of the review addresses cytoplasmic transport after escape. Viral particles exploit the microtubule network and motor proteins (kinesin, dynein) to achieve directed motion toward the nucleus. The authors construct a Langevin‑type model that couples motor stepping dynamics, binding/unbinding kinetics of nuclear localization signals (NLS), and the geometry of the microtubule meshwork. This active transport model predicts mean transit times on the order of minutes and a high probability of successful nuclear entry for small to medium‑sized viruses. In contrast, non‑viral polyplexes lack active propulsion and rely on passive diffusion through a crowded cytoplasm. To capture this, the authors introduce an effective diffusion coefficient that incorporates cytoplasmic viscosity, steric hindrance from organelles, and electrostatic interactions with the cytosolic environment. Simulations show that diffusion alone leads to transit times ranging from several hours to days, explaining the markedly lower transfection efficiencies of synthetic carriers.
Building on these two mechanistic frameworks, the review proposes a hybrid vector design strategy. By grafting viral fusogenic peptides or endosomolytic domains onto synthetic polymers, one can combine the efficient endosomal rupture of viruses with the safety and cargo capacity of non‑viral carriers. The authors integrate the proton‑spunge model with an active transport module that mimics motor‑driven movement, creating a multi‑scale simulation platform that explores a high‑dimensional parameter space (polymer charge density, peptide affinity, motor attachment probability, etc.). Optimization within this space identifies configurations that simultaneously maximize endosomal escape probability and minimize cytoplasmic transit time, offering concrete design guidelines for next‑generation gene delivery vectors.
Finally, the article acknowledges current limitations, such as cell‑type specific variations in endosomal pH regulation, the heterogeneous nature of the cytoplasmic crowding, and the scarcity of quantitative experimental data for model validation. The authors suggest future directions including coupling the models with live‑cell imaging data, employing machine‑learning techniques for parameter inference, and extending the framework to account for nuclear pore complex transport mechanisms. Overall, the review demonstrates that integrating stochastic Brownian dynamics, reaction‑diffusion kinetics, and microtubule‑based active transport into a unified modeling approach provides powerful predictive insight into the bottlenecks of gene delivery and guides rational engineering of more effective synthetic and hybrid viral vectors.
Comments & Academic Discussion
Loading comments...
Leave a Comment