Modelling Spatial Interactions in the Arbuscular Mycorrhizal Symbiosis using the Calculus of Wrapped Compartments
Arbuscular mycorrhiza (AM) is the most wide-spread plant-fungus symbiosis on earth. Investigating this kind of symbiosis is considered one of the most promising ways to develop methods to nurture plants in more natural manners, avoiding the complex chemical productions used nowadays to produce artificial fertilizers. In previous work we used the Calculus of Wrapped Compartments (CWC) to investigate different phases of the AM symbiosis. In this paper, we continue this line of research by modelling the colonisation of the plant root cells by the fungal hyphae spreading in the soil. This study requires the description of some spatial interaction. Although CWC has no explicit feature modelling a spatial geometry, the compartment labelling feature can be effectively exploited to define a discrete surface topology outlining the relevant sectors which determine the spatial properties of the system under consideration. Different situations and interesting spatial properties can be modelled and analysed in such a lightweight framework (which has not an explicit notion of geometry with coordinates and spatial metrics), thus exploiting the existing CWC simulation tool.
💡 Research Summary
**
The paper presents a novel application of the Calculus of Wrapped Compartments (CWC) to model spatial interactions in the arbuscular mycorrhizal (AM) symbiosis, focusing on the colonisation of plant root cells by fungal hyphae spreading through the soil. AM is a ubiquitous plant‑fungus partnership that contributes to nutrient uptake for the majority of terrestrial plants and is therefore a promising target for sustainable agriculture. While previous work by the authors employed CWC to describe molecular signalling and intracellular processes in AM, the spatial aspect of hyphal growth, diffusion of signalling molecules, and root penetration had not been addressed.
CWC Overview
CWC is a formal language built on an alphabet of atomic elements (atoms) and a set of compartment labels. A term consists of multisets of atoms and compartments; each compartment is defined by a wrap (membrane atoms), a content (inner term), and a label (type). System dynamics are expressed through rewrite rules of the form `: p → o, where p is a pattern and o an open term. Variables are restricted to a single wrap variable and a single content variable per compartment, guaranteeing unambiguous matching. By attaching a kinetic constant k to each rule, the authors obtain a stochastic semantics based on Gillespie’s direct method, allowing the calculation of reaction propensities as k multiplied by the number of possible reactant combinations. The existing CWC simulator implements this semantics, supports various kinetic laws, and runs parallel simulations on multi‑core architectures.
Spatial Modelling Strategy
CWC does not provide explicit coordinates, but the authors exploit the compartment label mechanism to create an implicit, discrete topology. Each spatial region—whether a soil patch, an epidermal cell, or a cortical cell—is represented by a uniquely labelled compartment. Adjacency relationships are encoded either by naming conventions (e.g., i,j ↔ i,j+1) or by explicit rewrite rules that allow interaction only between neighbouring labels. This approach eliminates the need for continuous distance calculations, collision detection, or geometric transformations, while still enabling the representation of diffusion, growth, and penetration processes.
Illustrative Examples
-
Cell Proliferation on a CWC Grid – The authors model a finite k × n lattice where each grid cell is a compartment labelled c. An atom e denotes an empty site, and a compartment with membrane label m and a single atom (M, G1, S, G2) represents a biological cell in a specific cell‑cycle phase. Rules allow a cell to move into an adjacent empty compartment and to progress through the cell‑cycle phases, demonstrating how spatial constraints can be imposed on population dynamics.
-
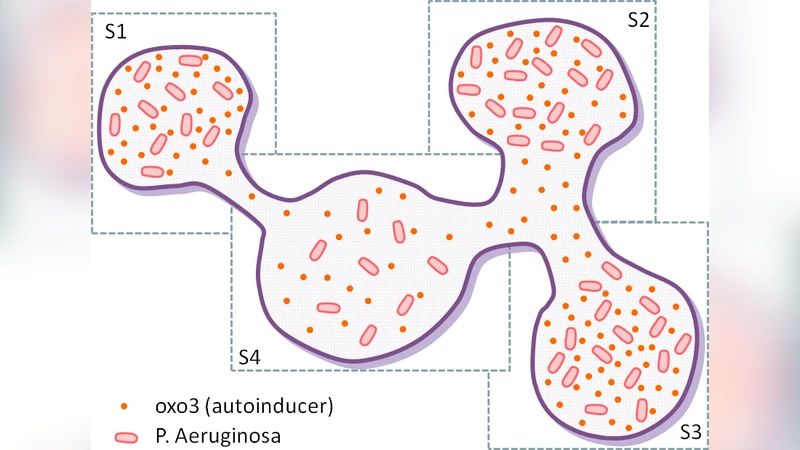
Quorum‑Sensing Diffusion – Four environmental sectors (S1…S4) are modelled as compartments with distinct labels. An auto‑inducer molecule (oxo3) diffuses between sectors with different rates, encoded by separate rules. Bacterial agents are compartments labelled m; when the local oxo3 concentration exceeds a threshold, the bacteria acquire an “active” label. Simulations show how heterogeneous diffusion capacities affect the timing and extent of quorum‑sensing activation, illustrating the ability of CWC to capture anisotropic spatial effects without explicit geometry.
AM Symbiosis Model
The core contribution is a CWC model of the AM colonisation process. The model includes:
-
Signal Molecule Diffusion – Plant roots release strigolactones (SL) into soil compartments; fungal hyphae respond by producing the Myc factor (MF). Both SL and MF are represented as atoms that move between adjacent soil compartments according to diffusion rules with rate constants reflecting experimental diffusion coefficients.
-
Hyphal Growth – Hyphae are modelled as the atom HY. A growth rule states that if a compartment contains HY, an adjacent empty soil compartment may receive a new HY atom, thereby extending the fungal network. The rule’s kinetic constant captures hyphal elongation speed.
-
Root Penetration – When a hyphal atom reaches a compartment labelled ROOT_EPI (epidermal cell), a penetration rule replaces the ROOT_EPI label with INFECTED_EPI and inserts a new compartment representing an intracellular hyphal segment.
-
Arbuscule Formation – Inside cortical cells (label ROOT_COR), a rule creates an ARBUSCULE compartment containing nutrient exchange atoms (e.g., P for phosphate, N for nitrogen). Subsequent exchange rules move these nutrients between the ARBUSCULE and the surrounding plant cytoplasm, modelling the bidirectional transfer that characterises functional mycorrhizae.
The authors executed thousands of stochastic simulations using the CWC simulator. Key observations include:
- A threshold concentration of strigolactone triggers a rapid increase in hyphal growth rate, reproducing experimentally observed “branching bursts.”
- The probability of successful root penetration correlates strongly with both the diffusion rate of signalling molecules and the hyphal elongation constant, highlighting the interplay between chemical and physical processes.
- Arbuscule formation occurs after a predictable delay once hyphae have entered cortical cells, matching temporal patterns reported in microscopy studies.
Discussion and Future Work
The paper argues that the label‑based topology offers a lightweight yet expressive means to encode spatial constraints, allowing the reuse of existing CWC tools without the overhead of geometric engines. Limitations are acknowledged: the approach cannot directly represent continuous distances, making quantitative calibration of diffusion coefficients to real‑world units non‑trivial; scaling to three‑dimensional tissues would require a combinatorial increase in compartment labels and rules. Future directions include hybrid models that combine discrete CWC compartments with explicit coordinate systems, hierarchical graph representations to reduce label explosion, and systematic parameter fitting against experimental time‑course data. The authors also envision extending the framework to other plant‑microbe interactions, such as rhizobial nodulation, thereby establishing a general computational platform for spatially resolved symbiosis modeling.
Comments & Academic Discussion
Loading comments...
Leave a Comment