Collective Cell Migration: Leadership, Invasion and Segregation
A number of biological processes, such as embryo development, cancer metastasis or wound healing, rely on cells moving in concert. The mechanisms leading to the emergence of coordinated motion remain however largely unexplored. Although biomolecular signalling is known to be involved in most occurrences of collective migration, the role of physical and mechanical interactions has only been recently investigated. In this paper, a versatile framework for cell motility is implemented in-silico in order to study the minimal requirements for the coordination of a group of epithelial cells. We find that cell motility and cell-cell mechanical interactions are sufficient to generate a broad array of behaviours commonly observed in vitro and in vivo. Cell streaming, sheet migration and susceptibility to leader cells are examples of behaviours spontaneously emerging from these simple assumptions, which might explain why collective effects are so ubiquitous in nature. This analysis provides also new insights into cancer metastasis and cell sorting, suggesting in particular that collective invasion might result from an emerging coordination in a system where single cells are mechanically unable to invade.
💡 Research Summary
The authors present a computational study of collective cell migration using a minimal, physics‑based framework that incorporates only two essential ingredients: a self‑propelled motile force (μ) generated by each cell and a mechanical coupling between neighboring cells represented by a membrane tension (J). Cells are modeled on a two‑dimensional lattice with the Cellular Potts Model (CPM). Each cell possesses a polarity vector n_i that determines the direction of its active force. Polarity evolves with a memory time τ, aligning gradually toward the cell’s recent displacement, thereby providing a feedback loop between motion and orientation. No explicit biochemical signaling is included; interactions are limited to volume exclusion, adhesion (through J), and the active force μ.
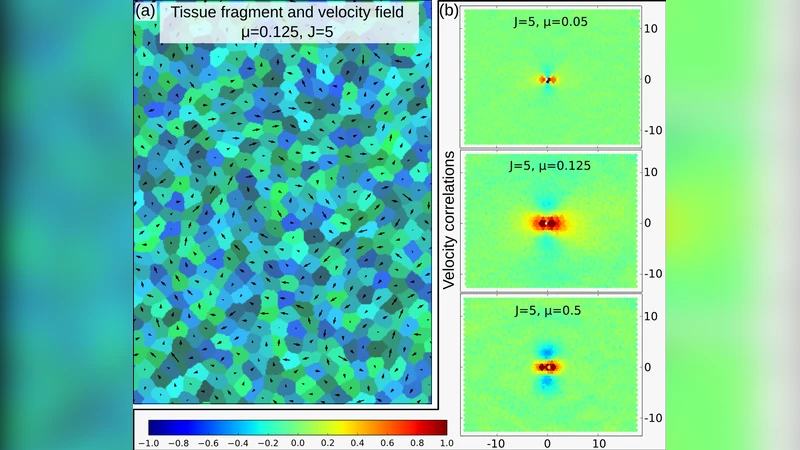
The study first explores how varying μ at fixed J influences the emergent dynamics of a dense epithelial sheet. At low μ the tissue remains static, with cells maintaining neighbor relationships. As μ surpasses a critical value μ_c (which scales with J), cells begin to stream: velocity correlation maps reveal long‑range alignment along the flow direction and anti‑alignment perpendicular to it, reproducing experimentally observed streaming patterns in endothelial monolayers. The authors quantify motion using the mean‑square displacement ⟨Δr²⟩ and fit it to a power law Δr² = DΔt^β. The exponent β serves as a dynamical indicator: β≈1 denotes normal diffusion, β<1 sub‑diffusion, and β>1 super‑diffusion. Near μ_c, β rises sharply above 1, indicating persistent, super‑diffusive motion consistent with coherent streams. At very high μ, β returns toward 1, reflecting that excessive propulsion disrupts coordination as cells penetrate each other, leading to essentially random walks.
Next, the authors examine system‑size effects. By simulating populations ranging from 9 to 1,600 cells while keeping density constant, they define a global order parameter h = ⟨v_i⟩/⟨|v_i|⟩ that measures the degree of collective alignment. For each (μ, J) pair a characteristic length λ_g(μ, J) emerges: if the linear size of the group d = √N is smaller than λ_g, the tissue spontaneously transitions from a static epithelium to a coordinated sheet that migrates as a unit. When d exceeds λ_g, only local streams persist and global order collapses. Thus, the ability of a cell population to achieve sheet‑like migration is intrinsically linked to the interplay between motility, adhesion, and group size.
The role of leader cells is probed by embedding a small fraction (≤5 %) of “leader” cells whose polarity is fixed externally rather than updated via the feedback rule. All leaders share the same fixed direction, and they are distributed uniformly throughout the tissue (not confined to edges). The key length scale becomes the average distance between leaders, d_l. When d_l < λ_g, the leader orientation propagates efficiently, raising the global order parameter to near‑unity; the tissue moves coherently toward the imposed direction. When d_l > λ_g, leader influence remains local and the overall tissue retains low alignment. This demonstrates that even without biochemical cues, a sparse set of mechanically distinct cells can steer an entire collective, provided their spacing is within the mechanically mediated communication range.
Finally, the authors explore invasion by increasing μ while decreasing J, mimicking a scenario where individual cells become mechanically capable of penetrating a surrounding matrix. In this regime, the collective loses its coherence and cells migrate independently, resembling a transition from collective to single‑cell invasion observed in certain cancers. The model thus captures an “EMT‑like” switch purely through mechanical parameters, suggesting that the epithelial‑mesenchymal transition may be driven, at least in part, by quantitative changes in cell motility and adhesion rather than by a binary genetic program.
Overall, the paper establishes that a simple self‑propelled, mechanically coupled cell model can reproduce a wide spectrum of collective behaviors—streaming, sheet migration, leader‑driven directionality, and invasion—without invoking complex biochemical networks. By systematically varying motile force, adhesion strength, group size, and leader density, the authors map out phase‑like transitions between distinct migratory modes and provide quantitative predictions (critical μ_c, λ_g, d_l) that can be tested experimentally. The work underscores the sufficiency of physical interactions in shaping tissue‑scale dynamics and offers a parsimonious framework for interpreting phenomena such as EMT, tumor invasion, and wound‑healing migration from a biophysical perspective.
Comments & Academic Discussion
Loading comments...
Leave a Comment