Direction dependent mechanical unfolding and Green Fluorescent Protein as a force sensor
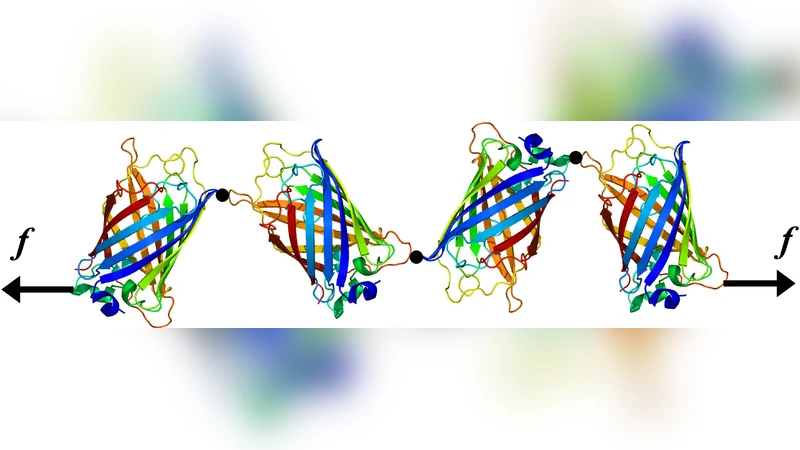
An Ising–like model of proteins is used to investigate the mechanical unfolding of the Green Fluorescent Protein along different directions. When the protein is pulled from its ends, we recover the major and minor unfolding pathways observed in experiments. Upon varying the pulling direction, we find the correct order of magnitude and ranking of the unfolding forces. Exploiting the direction dependence of the unfolding force at equilibrium, we propose a force sensor whose luminescence depends on the applied force.
💡 Research Summary
The paper presents a comprehensive study of the mechanical unfolding of Green Fluorescent Protein (GFP) using a native‑centric Ising‑like model, and it leverages the direction‑dependent unfolding forces to propose a GFP‑based molecular force sensor. The authors first adapt a previously developed Ising‑type model, in which each residue is described by a binary variable m_k indicating whether it is in a native‑like conformation, and each pair of residues i, j that belong to the same native stretch carries an orientation variable σ_ij = ±1. Native contacts contribute a negative interaction energy h_ij, and the total Hamiltonian consists of the sum of these contact terms plus a coupling term U(L) that accounts for the external pulling force. For a constant force, U = –f L; for a constant‑velocity protocol, a harmonic potential moving at speed v is used. Because the model is exactly solvable in equilibrium under constant force, the authors compute the free‑energy profile ΔG(L) as a function of the molecular extension L. At zero force the profile shows a native minimum, an unfolded minimum, and three intermediate minima around 11 nm, 18 nm, and 25 nm, which correspond to partial unfolding of specific β‑strands (β1/β11, β10/β11, and β1‑β3 respectively).
Dynamic Monte‑Carlo simulations are then performed to mimic AFM pulling experiments at constant velocities v = 0.3, 1, 2, and 3.6 µm s⁻¹. The simulations reproduce the two experimentally observed unfolding pathways: a major pathway (α‑helix → β1 → β2/β3 → β10/β11) occurring in roughly 70 % of trajectories, and a minor pathway (α‑helix → β11 → β10) in the remaining 30 %. The authors monitor the end‑to‑end length L, the instantaneous force, and the weighted fractions of native contacts φ_{βi‑βj} between adjacent β‑strands. The major pathway shows three distinct drops in φ corresponding to the sequential loss of contacts in β1, then β2‑β3, and finally β10‑β11, each associated with plateaus in L that match the intermediate minima of ΔG(L). The minor pathway displays an early loss of contacts in β11 followed by β10, consistent with experimental observations of barrel flattening and subsequent strand rupture.
To explore directionality, the authors pull not only from the protein ends but also from various internal residues, thereby changing the force vector relative to the β‑barrel. Table I reports the mean unfolding forces for several pulling directions (e.g., end‑end, 3‑212, 132‑212, 182‑212, 3‑132, 117‑182) at the three pulling speeds. Although the absolute forces are systematically higher (by a factor of 2–3) than experimental values, the ranking of forces across directions matches the experimental hierarchy: end‑end < 3‑212 ≈ 132‑212 < 182‑212 ≈ 3‑132 < 117‑182. This agreement demonstrates that the model captures the essential anisotropy of GFP’s mechanical stability despite its lack of full three‑dimensional detail.
The authors further fit the most probable rupture forces f_M versus pulling speed to the Evans–Ritchie expression f_M = (k_BT/x_u) ln(τ₀ x_u k_BT r), extracting an effective potential width x_u for each direction (Table II). The extracted x_u values agree with experiment only for a few pulling geometries, highlighting limitations of the one‑dimensional approximation and of the Evans–Ritchie assumption of a force‑independent barrier width.
Finally, the paper proposes a practical application: by assembling a poly‑protein composed of GFP modules linked such that each module experiences a different pulling direction, the overall construct can act as a force sensor. Because each GFP module loses fluorescence only when the applied force exceeds its direction‑specific rupture threshold, the pattern of fluorescence (which modules are bright or dark) directly encodes the magnitude of the applied force over a broad range. This concept overcomes the traditional limitation that GFP fluorescence is essentially binary (on/off) with respect to mechanical stress, enabling quantitative force read‑out in vivo.
In summary, the study validates an Ising‑like coarse‑grained model against detailed AFM data, elucidates the direction‑dependent mechanical unfolding landscape of GFP, and translates these insights into a novel design for a molecular force sensor based on GFP’s force‑sensitive fluorescence. The work bridges theoretical modeling, single‑molecule biophysics, and bio‑nanotechnological device design, offering a promising route toward real‑time, intracellular force measurement.
Comments & Academic Discussion
Loading comments...
Leave a Comment