Quantum Proteomics
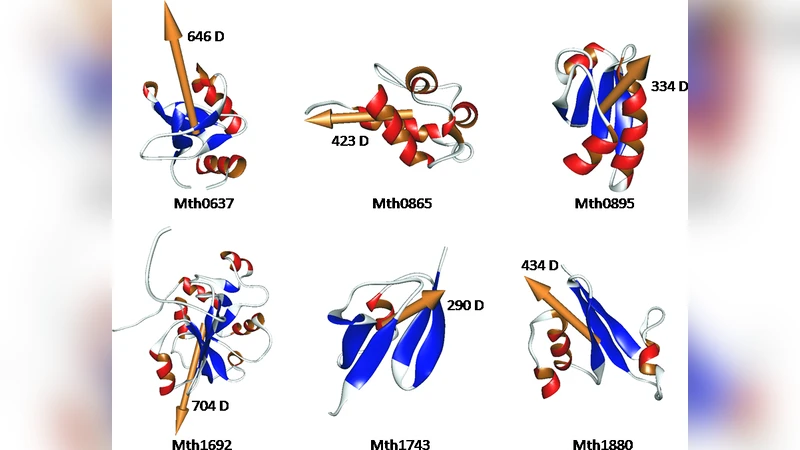
We put forward the idea of establishing a novel interdisciplinary field of research at the interface between quantum mechanics and proteomics. The new field, called quantum proteomics, is defined as the large-scale study of the electronic structure of the proteins that define an organism’s proteome. The electronic structure of proteins is unveiled with the aid of linear-scaling quantum mechanical calculations. Such calculations provide information about the energy levels of the proteins, the charges of their amino acid side chains, their electrostatic potentials and permanent dipole moments ({\mu}). Since the magnitude of the electric dipole moment of any protein is not null ({\mu}\neq0 Debye), the dipole moment can be employed to characterize the electronic structure of each protein that belongs to an organism’s proteome. As an example, we investigate six proteins from the thermophilic bacterium Methanobacterium thermoautotrophicum (Mth) whose atomic structures were characterized by solution NMR spectroscopy.
💡 Research Summary
The paper proposes a new interdisciplinary discipline called “quantum proteomics” (QP), positioned at the interface of proteomics and quantum mechanics. The authors define QP as the large‑scale study of the electronic structure of all proteins that constitute an organism’s proteome. To demonstrate feasibility, they performed linear‑scaling quantum mechanical (QM) calculations on six proteins from the thermophilic bacterium Methanobacterium thermoautotrophicum (Mth) whose three‑dimensional structures were determined by solution NMR. Calculations were carried out with the MOPAC2009 package using the MOZYME algorithm, which exploits a Lewis‑structure based initial guess to obtain localized orbitals and achieve near‑linear scaling even for systems containing several thousand atoms. The semi‑empirical PM6 Hamiltonian supplied the electronic Hamiltonian, while solvent effects were modeled with the COSMO continuum (dielectric constant ε = 78.4).
The QM results include the set of occupied molecular orbitals, atomic partial charges, electrostatic potentials, and, most importantly, the permanent electric dipole moment (μ) of each protein. Because any neutral or charged protein necessarily possesses a non‑zero dipole (μ ≠ 0 Debye), the authors argue that μ provides a compact, physically meaningful descriptor of a protein’s electronic structure. The six Mth proteins vary in size (1,120–3,219 atoms) and net charge (+3, 0, –2, –4, etc.), yet all exhibit sizable dipoles ranging from ~290 Debye to ~704 Debye. The authors illustrate the dipole vectors graphically, placing the tail at the protein’s center of mass and the tip indicating the positive pole. They note that the dipole arises from the spatial distribution of charged residues and the intrinsic peptide bond dipoles (~3.5 Debye each), and that surrounding counter‑ions have only a minor effect on the overall μ.
The central thesis is that dipole moments can serve as a practical index for classifying proteins within a proteome, complementing traditional sequence‑ or structure‑based approaches. By compiling dipole data (and other electronic properties such as energy levels) into a searchable database, researchers could correlate electronic signatures with biological functions, interaction partners, or enzymatic activities. The authors also suggest that coupling QM‑derived electronic information with classical molecular dynamics (MD) simulations would enable the study of dynamic dipole fluctuations and their role in recognition events.
Limitations are acknowledged: the PM6 semi‑empirical method, while efficient, is an approximation and may not capture subtle electronic effects, especially for metal‑containing sites or non‑standard residues. Using raw NMR models without further geometry optimization introduces structural uncertainty, and the choice of reference point for dipole calculation (center of mass vs. charge centroid) can affect the reported values. Consequently, standardized protocols and validation against higher‑level quantum methods are needed before large‑scale deployment.
In conclusion, the authors envision quantum proteomics as a foundational framework that bridges molecular electronic structure and functional proteomics. They anticipate that, as computational methods improve and comprehensive electronic‑structure databases become available, QP will provide new insights into protein‑protein interactions, signaling pathways, and enzymatic mechanisms, ultimately enriching our quantitative understanding of biological systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment