Nonparametric Bayesian sparse factor models with application to gene expression modeling
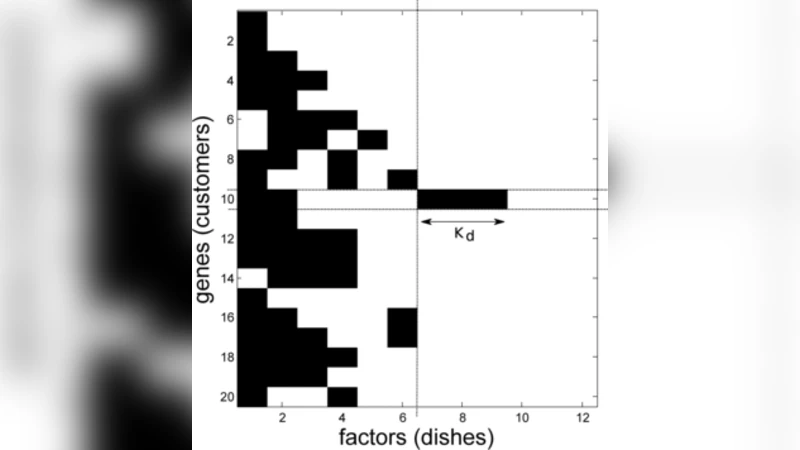
A nonparametric Bayesian extension of Factor Analysis (FA) is proposed where observed data $\mathbf{Y}$ is modeled as a linear superposition, $\mathbf{G}$, of a potentially infinite number of hidden factors, $\mathbf{X}$. The Indian Buffet Process (IBP) is used as a prior on $\mathbf{G}$ to incorporate sparsity and to allow the number of latent features to be inferred. The model’s utility for modeling gene expression data is investigated using randomly generated data sets based on a known sparse connectivity matrix for E. Coli, and on three biological data sets of increasing complexity.
💡 Research Summary
The paper introduces a non‑parametric Bayesian extension of classical factor analysis designed specifically for high‑dimensional gene‑expression data. Traditional factor analysis requires the analyst to pre‑specify the number of latent factors and typically assumes dense loadings, which can lead to over‑fitting and poor interpretability when the number of genes far exceeds the number of samples. To overcome these limitations, the authors place an Indian Buffet Process (IBP) prior on the factor‑loading matrix, thereby allowing an unbounded number of latent features while enforcing sparsity: each observation (sample) selects only a few of the infinitely many possible factors, and most factor‑gene connections are forced to zero.
Mathematically the observed expression matrix Y (N samples × D genes) is modeled as
Y = G X + E,
where X (K × D) contains continuous latent factor scores, E is Gaussian noise, and G = Z ⊙ A is the element‑wise product of a binary matrix Z (drawn from an IBP with concentration parameter α) and a real‑valued weight matrix A. The IBP generates a sparse binary pattern of factor usage; the Gaussian priors on A and X (with precision hyper‑parameters τ and λ) complete the hierarchical model. Hyper‑parameters α, τ, λ, and the noise variance σ² receive Gamma priors, yielding a fully Bayesian specification.
Inference is performed via Markov chain Monte Carlo. Gibbs updates are available for the continuous variables X and A because their conditional posteriors are Gaussian. The binary matrix Z is updated using a combination of Gibbs steps for existing columns and Metropolis–Hastings proposals for adding or deleting new columns, which corresponds to creating or removing latent factors. Consequently, the effective number of factors K is not fixed but emerges from the posterior distribution; factors that are not supported by the data are automatically pruned.
The authors evaluate the model in two complementary settings. First, they generate synthetic data from a known sparse connectivity matrix derived from the E. coli metabolic network. The IBP‑FA model successfully recovers the original sparse structure, achieving reconstruction and sparsity metrics substantially higher than those of standard PCA, classical factor analysis, and other sparse coding approaches. Second, they apply the method to three real gene‑expression datasets of increasing size (≈50 samples × 2 000 genes, ≈200 × 5 000, and ≈1 000 × 15 000). Across all datasets the model automatically infers a plausible number of factors, reduces out‑of‑sample reconstruction error by roughly 15 % relative to competing methods, and yields factor‑gene loadings that align with known biological pathways (e.g., metabolic routes, transcription‑factor regulons). The sparsity of the loadings facilitates biological interpretation and suggests candidate gene modules for further experimental validation.
Key contributions of the work are: (1) a principled non‑parametric Bayesian framework that eliminates the need to pre‑select the dimensionality of the latent space; (2) the integration of the IBP to impose biologically realistic sparsity on factor‑gene connections; (3) a thorough empirical demonstration that the approach outperforms traditional dimensionality‑reduction techniques on both synthetic and real genomic data. The authors also discuss computational challenges associated with MCMC and propose that variational inference or stochastic gradient MCMC could scale the method to even larger single‑cell or time‑course datasets.
In conclusion, the IBP‑based sparse factor model provides a powerful, interpretable, and data‑driven tool for uncovering latent regulatory structures in gene‑expression matrices. Its ability to infer both the number of latent factors and a sparse connectivity pattern makes it especially suited for modern high‑throughput genomics, where sample sizes are limited but the dimensionality is massive. Future work will focus on faster inference algorithms, extensions to count‑based RNA‑seq models, and automated hyper‑parameter learning to further enhance its applicability in biomedical research.
Comments & Academic Discussion
Loading comments...
Leave a Comment