Protein Models Comparator: Scalable Bioinformatics Computing on the Google App Engine Platform
The comparison of computer generated protein structural models is an important element of protein structure prediction. It has many uses including model quality evaluation, selection of the final models from a large set of candidates or optimisation of parameters of energy functions used in template-free modelling and refinement. Although many protein comparison methods are available online on numerous web servers, they are not well suited for large scale model comparison: (1) they operate with methods designed to compare actual proteins, not the models of the same protein, (2) majority of them offer only a single pairwise structural comparison and are unable to scale up to a required order of thousands of comparisons. To bridge the gap between the protein and model structure comparison we have developed the Protein Models Comparator (pm-cmp). To be able to deliver the scalability on demand and handle large comparison experiments the pm-cmp was implemented “in the cloud”. Protein Models Comparator is a scalable web application for a fast distributed comparison of protein models with RMSD, GDT TS, TM-score and Q-score measures. It runs on the Google App Engine (GAE) cloud platform and is a showcase of how the emerging PaaS (Platform as a Service) technology could be used to simplify the development of scalable bioinformatics services. The functionality of pm-cmp is accessible through API which allows a full automation of the experiment submission and results retrieval. Protein Models Comparator is free software released on the Affero GNU Public Licence and is available with its source code at: http://www.infobiotics.org/pm-cmp This article presents a new web application addressing the need for a large-scale model-specific protein structure comparison and provides an insight into the GAE (Google App Engine) platform and its usefulness in scientific computing.
💡 Research Summary
The paper presents Protein Models Comparator (pm‑cmp), a web‑based service designed to perform large‑scale comparisons of protein structural models, a task that is central to protein structure prediction pipelines. Traditional online structure comparison tools are geared toward comparing experimentally determined proteins, typically offering only single pairwise (1:1) comparisons and lacking the ability to handle thousands of model‑to‑model comparisons required in modern prediction experiments such as CASP. To fill this gap, the authors built pm‑cmp on Google App Engine (GAE), a Platform‑as‑a‑Service (PaaS) cloud environment that provides automatic scaling of compute resources without the need for users to manage hardware.
Key technical contributions include:
-
Model‑specific comparison logic – Because the compared entities are models of the same protein, the atomic correspondence is known a priori. pm‑cmp extracts the common Cα atoms from each pair of models, thereby fixing the alignment and allowing the use of standard similarity measures even when models are incomplete. This “matching residues” step is not an alignment in the classical sense but a preprocessing step that selects the intersecting set of residues, handling cases where residues are missing at the N‑ or C‑termini or where templates do not cover the full sequence.
-
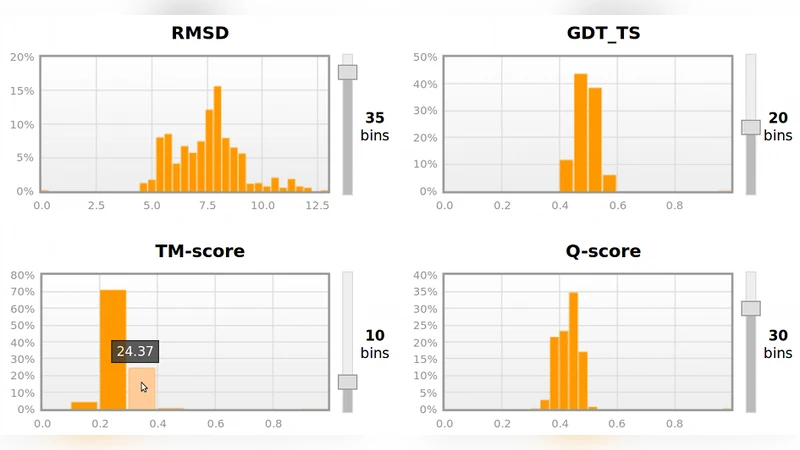
Support for multiple similarity metrics – The service implements four widely used scores: RMSD, GDT‑TS, TM‑score, and Q‑score. For GDT‑TS and TM‑score the user can choose between scaling by the number of matched residues or by the total length of the structures, providing flexibility for evaluating partial models versus full‑length models.
-
Scalable cloud architecture – The front‑end UI is written in Python using the Web2py framework and runs in the GAE Python runtime. The heavy‑weight comparison engine is implemented in Groovy on top of the Gaelyk lightweight Java framework, leveraging the BioShell library for the actual structural calculations. Communication between UI and engine occurs via HTTP requests.
-
Task queue based parallelisation – Upon experiment launch, each structure‑versus‑structure comparison is encapsulated as an individual task and placed into GAE’s Task Queue. A token‑bucket algorithm controls the maximum number of concurrent tasks, while GAE automatically spawns additional application instances to satisfy demand. Results are stored in the Datastore, and repeated reads of the same structure are served from memcache to minimise latency.
-
RESTful API for full automation – The service exposes endpoints for experiment setup, model upload, experiment start, status polling, and result download. This enables programmatic submission of thousands of models, integration into pipelines (e.g., clustering, energy‑function optimisation), and batch processing without manual interaction.
Performance testing demonstrated that a 1 : N experiment with 5 000 models (≈150 MB of data) completed in under 12 minutes, with total cost staying within GAE’s free tier (daily limits on CPU, storage, and bandwidth). The authors discuss platform limitations: GAE enforces a 30‑second request timeout, a 1 GB per‑instance memory cap, and restrictions on using native C/C++ libraries in the Java runtime. They propose future migration to Docker‑based Cloud Run or Kubernetes to overcome these constraints, allowing longer‑running jobs, larger memory footprints, and the inclusion of additional metrics such as lDDT or CAD‑score.
In summary, pm‑cmp demonstrates how a modern PaaS cloud can be harnessed to deliver a robust, scalable, and cost‑effective solution for high‑throughput protein model comparison, addressing a clear need in the structural bioinformatics community and offering a blueprint for similar large‑scale scientific computing services.
Comments & Academic Discussion
Loading comments...
Leave a Comment