Higher Order Programming to Mine Knowledge for a Modern Medical Expert System
Knowledge mining is the process of deriving new and useful knowledge from vast volumes of data and background knowledge. Modern healthcare organizations regularly generate huge amount of electronic data stored in the databases. These data are a valuable resource for mining useful knowledge to help medical practitioners making appropriate and accurate decision on the diagnosis and treatment of diseases. In this paper, we propose the design of a novel medical expert system based on a logic-programming framework. The proposed system includes a knowledge-mining component as a repertoire of tools for discovering useful knowledge. The implementation of classification and association mining tools based on the higher order and meta-level programming schemes using Prolog has been presented to express the power of logic-based language. Such language also provides a pattern matching facility, which is an essential function for the development of knowledge-intensive tasks. Besides the major goal of medical decision support, the knowledge discovered by our logic-based knowledge-mining component can also be deployed as background knowledge to pre-treatment data from other sources as well as to guard the data repositories against constraint violation. A framework for knowledge deployment is also presented.
💡 Research Summary
The paper addresses the growing need to extract useful knowledge from the massive electronic health records generated by modern healthcare organizations and to embed that knowledge into a decision‑support expert system. The authors propose a novel medical expert system built on a logic‑programming framework, specifically using SWI‑Prolog, and they focus on the implementation of two core knowledge‑mining components: a classification engine based on the ID3 decision‑tree algorithm and an association‑rule mining engine based on the Apriori algorithm.
Both engines are implemented using higher‑order and meta‑level programming techniques available in Prolog. For classification, the system dynamically asserts “node/2” and “edge/3” predicates to represent the decision tree, where each node stores lists of positive and negative case identifiers. The algorithm iteratively computes the information gain (Info) for each candidate attribute, selects the attribute with the minimum Info, and recursively splits the data until pure leaves are obtained. The resulting tree can be queried directly in Prolog, and the final classification rule is expressed as a Horn clause (e.g., class(allergy) :- fever=no, swollenGlands=no).
The association‑mining component leverages higher‑order predicates such as maplist, include, and setof to concisely generate candidate itemsets, count their support, and prune infrequent candidates. The implementation follows the classic Apriori approach: it first creates 1‑itemsets, then iteratively expands them while respecting a user‑defined minimum support threshold. Frequent patterns are output as sets of co‑occurring items, which can be transformed into association rules (e.g., fever=yes → class=no).
The overall system architecture consists of five layers: (1) a data‑integration module that aggregates heterogeneous clinical data (including textual documents) and stores it in a data warehouse; (2) the knowledge‑mining module described above; (3) a knowledge base that holds both induced rules and expert‑encoded background knowledge; (4) an inference and reasoning engine that interacts with medical practitioners; and (5) an OLAP interface for ad‑hoc querying. The authors illustrate the workflow with a small allergy‑diagnosis dataset containing ten patient records. Running the classification engine produces a decision tree whose leaf node isolates the allergy cases; running the association engine with a 50 % support threshold yields six frequent patterns, each interpretable as a diagnostic rule.
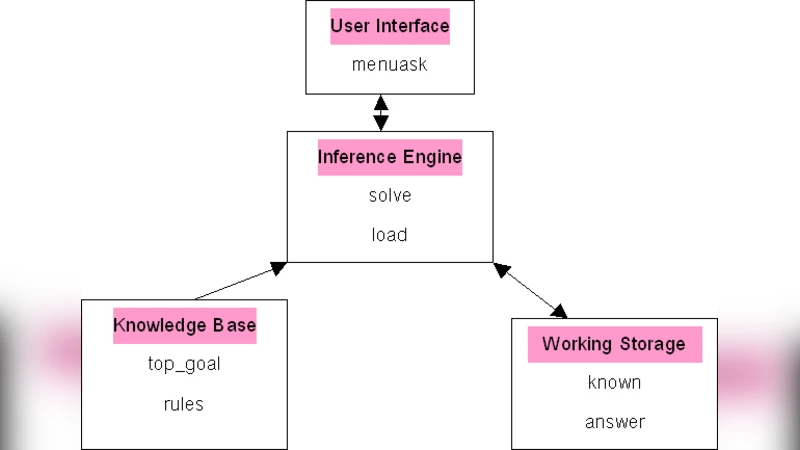
A key contribution is the “knowledge deployment” strategy. The induced rules can be automatically transformed into database triggers that enforce integrity constraints: any attempted insertion that violates a rule generates an error, thereby preventing inconsistent data. Additionally, the same rules can be loaded into an expert‑system shell; users can query the system and request explanations (“why?”), which are answered by the Horn clauses derived from the mining process. Figures in the paper depict the trigger generation pipeline, the automatically built knowledge base, the expert‑system shell architecture, and a sample command‑line interaction.
In the related‑work section, the authors compare their approach with other medical data‑mining efforts that rely on statistical or machine‑learning techniques, emphasizing the declarative nature and pattern‑matching strengths of logic programming. The paper concludes that higher‑order Prolog provides a concise, extensible platform for building knowledge‑intensive medical systems, but acknowledges that the experimental evaluation is limited to a toy dataset. Future work is suggested in scaling the system to larger clinical databases, conducting quantitative performance analyses, and automating the expert validation step for induced rules.
Overall, the paper demonstrates a proof‑of‑concept that integrates logic‑programming‑based knowledge discovery with a medical expert system, showing how mined rules can be reused both for decision support and for safeguarding data integrity.
Comments & Academic Discussion
Loading comments...
Leave a Comment