Computing fluxes and chemical potential distributions in biochemical networks: energy balance analysis of the human red blood cell
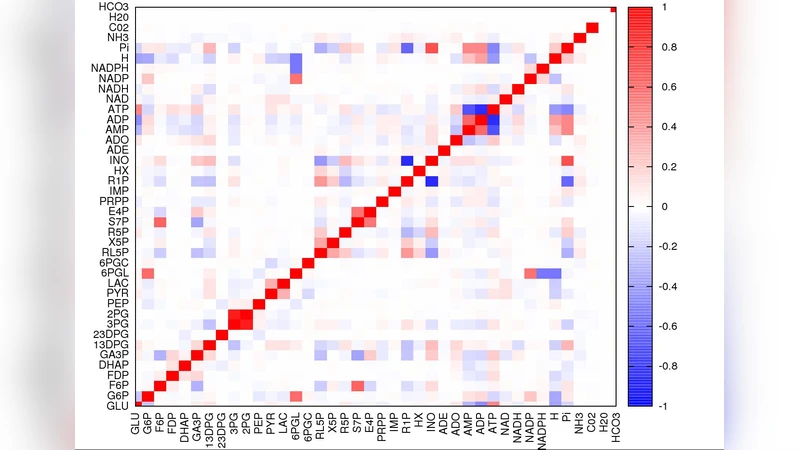
The analysis of non-equilibrium steady states of biochemical reaction networks relies on finding the configurations of fluxes and chemical potentials satisfying stoichiometric (mass balance) and thermodynamic (energy balance) constraints. Efficient methods to explore such states are crucial to predict reaction directionality, calculate physiologic ranges of variability, estimate correlations, and reconstruct the overall energy balance of the network from the underlying molecular processes. While different techniques for sampling the space generated by mass balance constraints are currently available, thermodynamics is generically harder to incorporate. Here we introduce a method to sample the free energy landscape of a reaction network at steady state. In its most general form, it allows to calculate distributions of fluxes and concentrations starting from trial functions that may contain prior biochemical information. We apply our method to the human red blood cell’s metabolic network, whose space of mass-balanced flux states has been sampled extensively in recent years. Specifically, we profile its thermodynamically feasible flux configurations, characterizing in detail how fluctuations of fluxes and potentials are correlated. Based on this, we derive the cell’s energy balance in terms of entropy production, chemical work done and thermodynamic efficiency.
💡 Research Summary
The paper addresses a central challenge in systems biology: how to characterize the set of feasible steady‑state configurations of a biochemical reaction network when both mass‑balance (stoichiometric) and thermodynamic (energy‑balance) constraints must be satisfied. While many tools exist for sampling the solution space defined solely by mass‑balance (e.g., Flux Balance Analysis, Monte‑Carlo sampling, message‑passing), incorporating the second law of thermodynamics remains difficult because reaction directions must be consistent with the sign of the Gibbs free‑energy change for each reaction.
To overcome this, the authors introduce two complementary algorithms based on the perceptron‑learning “MinOver” scheme. Method (a) treats the flux and free‑energy problems sequentially. First, a set of flux vectors that satisfy the stoichiometric constraints (either the exact mass‑balance equation Ξν = 0 or the more permissive Von Neumann production constraint Ξν ≥ 0) is sampled using existing Monte‑Carlo or MinOver techniques. The sign vector s = sign(ν) obtained from these fluxes is then used as a target for a second MinOver loop that adjusts the chemical potentials g = (gμ) so that the thermodynamic inequalities s_i ΔG_i ≤ 0 hold for all reactions. The update rule g_μ ← g_μ − α s_i ξ_μi (where ξ_μi are the stoichiometric coefficients) iteratively reduces the most violated inequality, guaranteeing convergence for any positive learning rate α. By repeatedly drawing initial g from a trial distribution P₀(g) that may encode experimental estimates, the algorithm produces a correlated ensemble of chemical‑potential vectors consistent with the sampled flux directions.
Method (b) removes the need for any a priori reversibility assignment and solves the flux and free‑energy problems jointly. All reactions are initially assumed reversible. Starting from a trial distribution of potentials, reaction affinities ΔG_i are computed and reaction directions s_i are set to the sign that makes the affinity downhill (s_i = −sign(ΔG_i)). The absolute flux magnitudes φ_i are drawn from a uniform distribution. The algorithm then enforces the Von Neumann production constraint in a softened form parameterized by ρ ∈
Comments & Academic Discussion
Loading comments...
Leave a Comment