Simulation in Statistics
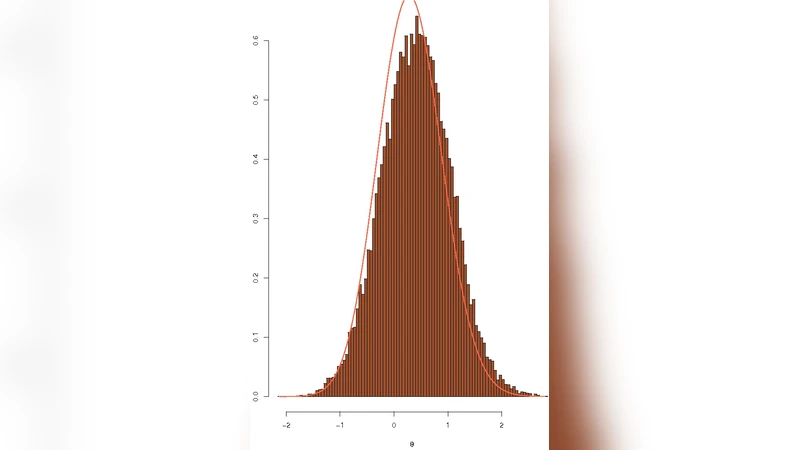
Simulation has become a standard tool in statistics because it may be the only tool available for analysing some classes of probabilistic models. We review in this paper simulation tools that have been specifically derived to address statistical challenges and, in particular, recent advances in the areas of adaptive Markov chain Monte Carlo (MCMC) algorithms, and approximate Bayesian calculation (ABC) algorithms.
💡 Research Summary
The paper “Simulation in Statistics” provides a comprehensive review of simulation techniques that have become indispensable in modern statistical practice. It begins by highlighting the natural synergy between statistics and simulation: statistical inference is fundamentally probabilistic, and stochastic simulation offers a flexible way to explore complex probabilistic models that are analytically intractable. The authors trace the historical roots of simulation, noting early mechanical devices such as Galton’s quincunx, Fisher’s randomised experiments, and Efron’s bootstrap, all of which foreshadowed the computational revolution that would later be driven by Monte Carlo methods.
Section 2 focuses on Monte Carlo methods in statistics. The authors discuss three major statistical paradigms where simulation is essential. First, the bootstrap is presented as a method that replaces analytical derivations of sampling distributions with empirical resampling from the observed data. The bootstrap’s reliance on the empirical cumulative distribution function (ECDF) makes it inherently a simulation technique. Second, maximum likelihood estimation (MLE) is examined, especially in contexts where the likelihood is multimodal (e.g., mixture models) or involves latent variables (e.g., stochastic volatility models). In such cases, closed‑form solutions are unavailable, and Monte Carlo integration or importance sampling become the tools of choice. Third, Bayesian inference is explored; the posterior distribution and Bayes factors typically involve high‑dimensional integrals that cannot be evaluated analytically. The authors illustrate this with a generalized linear model where testing a regression coefficient requires integrating over a high‑dimensional parameter space.
Section 3 introduces Markov chain Monte Carlo (MCMC) algorithms, the workhorse of modern Bayesian computation. The Metropolis–Hastings algorithm and Gibbs sampler are described in detail, emphasizing the detailed‑balance condition that guarantees the target distribution as the stationary distribution of the chain. A historical note points out the surge in “posterior distribution” mentions after Gelfand and Smith’s 1990 Gibbs sampler paper, underscoring MCMC’s impact on the field. The authors then discuss practical challenges: choosing an appropriate proposal distribution, scaling in high dimensions, and navigating multimodal landscapes. They argue that a one‑size‑fits‑all MCMC sampler is impossible because the very complexity of the target distribution motivates the use of MCMC.
Consequently, the paper delves into adaptive MCMC. It explains why early iterations of a chain can provide valuable information about the target’s geometry, suggesting that this information should be used to tune the proposal distribution on the fly. However, adaptation destroys the Markov property, invalidating classical ergodic theorems. The authors review seminal contributions that restore theoretical guarantees: regeneration‑based block independence (Gilks, Roberts, Sahu 1998), covariance‑adaptation schemes (Haario, Saksman, Tamminen 1999, 2001), and the general adaptive framework of Andrieu and Robert (2001). An illustrative example with a t‑distribution shows that naïve continual adaptation can introduce bias, reinforcing the need for carefully designed adaptation schedules (e.g., stopping adaptation after burn‑in).
Section 4 turns to Approximate Bayesian Computation (ABC), a set of methods developed to handle models with intractable likelihoods, originally in population genetics. ABC replaces likelihood evaluation with a simulation‑based acceptance step: parameters are drawn from the prior, synthetic data are generated, and the distance between summary statistics of synthetic and observed data is compared to a tolerance ε. The authors discuss the critical choices of summary statistics, distance metrics, and tolerance levels, and they note recent advances that embed ABC within Sequential Monte Carlo (SMC) frameworks, dramatically improving efficiency and allowing for adaptive tolerance reduction.
In the concluding remarks, the authors synthesize the narrative: simulation has evolved from a curiosity to a cornerstone of statistical methodology. Bootstrap and MLE illustrate how simulation resolves problems that lack analytical solutions. MCMC extends this capability to high‑dimensional Bayesian inference, while adaptive MCMC and ABC represent the frontier, addressing proposal tuning and likelihood intractability, respectively. The paper underscores that ongoing research in adaptive algorithms, variance reduction, and scalable ABC will continue to shape the future of statistical computation.
Comments & Academic Discussion
Loading comments...
Leave a Comment