Stochastic approximation to the specific response of the immune system
We develop a stochastic model to study the specific response of the immune system. The model is based on the dynamical interaction between Regulatory and Effector CD4+ T cells in the presence of Antigen Presenting Cells inside a lymphatic node. At a mean field level the model predicts the existence of different regimes where active, tolerant, or cyclic immune responses are possible. To study the model beyond mean field and to understand the specific responses of the immune system we use the Linear Noise Approximation and show that fluctuations due to finite size effects may strongly alter the mean field scenario. Moreover, it was found the existence of a certain characteristic frequency for the fluctuations. All the analytical predictions were compared with simulations using the Gillespie’s algorithm.
💡 Research Summary
This paper presents a stochastic framework for modeling the specific immune response mediated by the interaction of regulatory (R) and effector (E) CD4⁺ T cells in the presence of antigen‑presenting cells (APCs) within a lymph node. The authors begin by highlighting the limitations of traditional mean‑field approaches, which treat the lymph node as a homogeneous population of ~10⁷ lymphocytes, whereas individual T‑cell clones that mount a specific response consist of only 10²–10⁴ cells. Such small numbers render deterministic differential‑equation models inadequate for capturing clonal dynamics.
To overcome this, the authors first formulate the full master equation governing the probability distribution of the system’s state vector (numbers of free and APC‑bound R and E cells). Recognizing that exact solutions are rarely tractable, they adopt two complementary strategies: (i) stochastic simulation via the Gillespie algorithm, which provides exact sample paths consistent with the master equation, and (ii) analytical approximation using the Linear Noise Approximation (LNA). In the LNA, each population variable n_i is decomposed into a deterministic concentration x_i (scaling with the system size Ω_i) plus a fluctuation term proportional to √Ω_i β_i. Expanding the master equation in powers of Ω_i yields, at leading order, the deterministic rate equations (the mean‑field ODEs) and, at next order, a Fokker‑Planck equation that is equivalent to a set of linear Langevin equations with drift matrix A (the Jacobian of the deterministic system) and diffusion matrix D (determined by the reaction stoichiometry and propensities). The stationary covariance matrix Ξ satisfies the Lyapunov equation A Ξ + Ξ Aᵀ + D = 0, guaranteeing Gaussian fluctuations around any fixed point. By Fourier transforming the Langevin equations, the authors obtain explicit expressions for the power spectra P_i(ω) of each species, revealing how different frequencies contribute to the stochastic oscillations.
Beyond the LNA, the paper introduces the Effective Stability Approximation (ESA), which quantifies how intrinsic noise modifies the eigenvalues λ_i of the Jacobian. The correction term λ_i^corr scales as 1/Ω, so for small clone sizes the sign of λ_i can change, turning a mean‑field stable fixed point into an unstable one (or vice versa). This insight explains why fluctuations can induce transitions that are invisible in deterministic analyses.
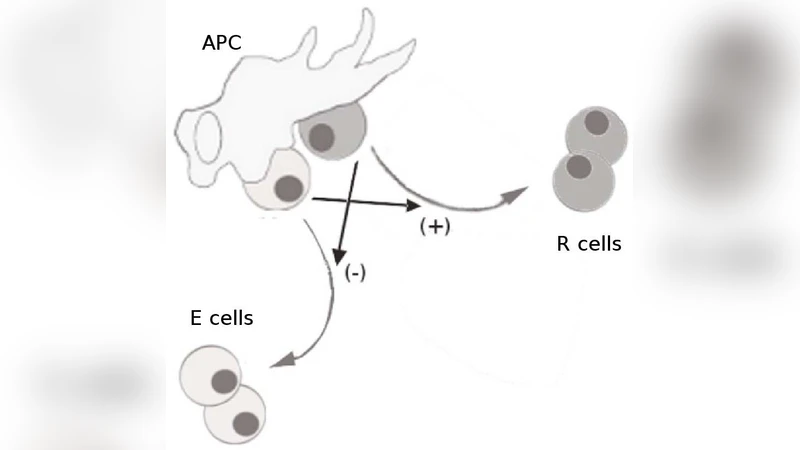
The biological model itself is grounded in experimental observations: (1) an effector T cell can proliferate only when bound to an APC that is not already occupied by a regulatory cell; (2) a regulatory T cell can proliferate only when bound to an APC that also hosts at least one effector cell. These rules generate six elementary reactions (binding/unbinding and division of each cell type) with rates that depend on the number of available APC binding sites. The resulting deterministic system exhibits three qualitatively distinct regimes: an “active” state with high effector numbers, a “tolerant” state dominated by regulators, and a limit‑cycle regime where populations oscillate.
Applying the LNA to each regime, the authors find that finite‑size fluctuations can dramatically reshape the phase diagram. In the tolerant regime, noise can push the system into the active basin, while in the limit‑cycle region the stochastic power spectrum displays a pronounced peak at a characteristic frequency, indicating noise‑enhanced oscillations. ESA calculations confirm that the effective eigenvalues shift enough to alter stability for realistic clone sizes (Ω≈10³).
To validate the analytical predictions, extensive Gillespie simulations were performed across a range of parameters and system sizes. The simulations reproduce the LNA‑predicted variance, correlation times, and spectral peaks, and they also reveal noise‑induced transitions not captured by deterministic equations. For example, trajectories that would settle into a tolerant fixed point in the mean‑field model occasionally escape to sustained oscillations due to stochastic kicks.
In conclusion, the study demonstrates that intrinsic demographic noise is a decisive factor in shaping specific immune responses at the clonal level. The combination of LNA, ESA, and exact stochastic simulation provides a powerful toolkit for dissecting complex immunological networks where cell numbers are low. The identification of a characteristic fluctuation frequency suggests potential avenues for diagnostic or therapeutic exploitation, such as targeting the timing of interventions to coincide with or disrupt these intrinsic oscillations. Future extensions could incorporate heterogeneous APC populations, cytokine‐mediated signaling, or spatial compartmentalization to further bridge the gap between theoretical models and experimental immunology.
Comments & Academic Discussion
Loading comments...
Leave a Comment