Simulating Spiking Neural P systems without delays using GPUs
We present in this paper our work regarding simulating a type of P system known as a spiking neural P system (SNP system) using graphics processing units (GPUs). GPUs, because of their architectural optimization for parallel computations, are well-suited for highly parallelizable problems. Due to the advent of general purpose GPU computing in recent years, GPUs are not limited to graphics and video processing alone, but include computationally intensive scientific and mathematical applications as well. Moreover P systems, including SNP systems, are inherently and maximally parallel computing models whose inspirations are taken from the functioning and dynamics of a living cell. In particular, SNP systems try to give a modest but formal representation of a special type of cell known as the neuron and their interactions with one another. The nature of SNP systems allowed their representation as matrices, which is a crucial step in simulating them on highly parallel devices such as GPUs. The highly parallel nature of SNP systems necessitate the use of hardware intended for parallel computations. The simulation algorithms, design considerations, and implementation are presented. Finally, simulation results, observations, and analyses using an SNP system that generates all numbers in $\mathbb N$ - {1} are discussed, as well as recommendations for future work.
💡 Research Summary
The paper presents a method for simulating spiking neural P systems (SNP systems) without delays on graphics processing units (GPUs) using the CUDA programming model. SNP systems are a class of membrane computing models that mimic the behavior of biological neurons: they consist of a single object alphabet {a}, a set of neurons each containing an initial number of spikes, and a finite set of firing rules. The authors focus on the variant where spikes are emitted immediately (no delay), which simplifies the operational semantics and makes the system amenable to matrix representation.
A key contribution is the formal translation of an SNP system into linear algebraic form. Each firing rule is represented as a row in a transition matrix M, while each neuron corresponds to a column. The current configuration of spikes across all neurons is stored in a vector C_k, and a binary “spiking vector” S_k indicates which rules fire at step k. The evolution of the system is captured by the equation C_{k+1} = C_k + S_k·M. This compact formulation enables the use of dense matrix‑vector multiplication to compute the next configuration in a single parallel operation.
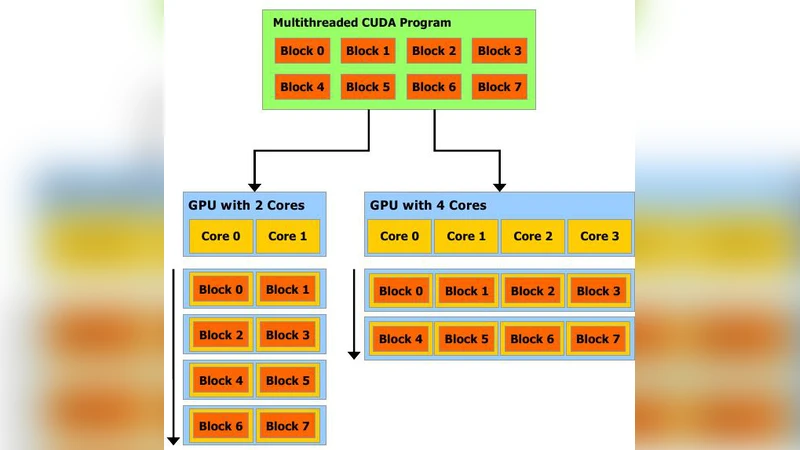
The implementation leverages NVIDIA’s CUDA architecture. Input data (the matrix M, the initial configuration C_0, and the set of admissible spiking vectors Ψ) are stored in plain text files and loaded on the host CPU. After allocating device memory, the data are transferred to the GPU using cudaMalloc and cudaMemcpy. The core kernel is launched with one thread per rule («<N,1»> where N is the number of rules). Each thread reads the corresponding row of M, multiplies it by the binary flag from S_k, and atomically adds the result to a temporary vector. After the kernel finishes, the resulting configuration vector is copied back to the host, where validity of the next spiking vector is checked (to enforce the non‑deterministic choice of a single applicable rule per neuron). This host‑side validation reduces branching inside the GPU kernel and improves overall throughput.
Experimental evaluation uses a well‑known SNP system that generates all natural numbers except 1, which is known to be non‑terminating and to produce an unbounded sequence of spikes. The authors compare execution times on a modern NVIDIA GPU against a sequential CPU implementation. Results show a speed‑up of several times, with performance scaling roughly linearly with the number of rules, confirming that the matrix‑based approach exploits the data‑parallel nature of GPUs effectively. The paper also discusses limitations: the current implementation stores M as a dense matrix, which can be memory‑inefficient for large, sparsely connected neural networks. The authors suggest future work on sparse matrix formats (e.g., CSR), multi‑GPU distribution, and extending the simulator to handle delayed SNP systems.
In conclusion, the study demonstrates that SNP systems, despite being theoretically powerful (Turing‑complete), can be simulated efficiently on commodity parallel hardware by casting their dynamics into linear algebra operations. The presented CUDA‑based simulator provides a foundation for large‑scale experiments with biologically inspired computing models and opens avenues for further optimization and extension to more complex variants of membrane computing.
Comments & Academic Discussion
Loading comments...
Leave a Comment