Finding Exogenous Variables in Data with Many More Variables than Observations
Many statistical methods have been proposed to estimate causal models in classical situations with fewer variables than observations (p<n, p: the number of variables and n: the number of observations). However, modern datasets including gene expression data need high-dimensional causal modeling in challenging situations with orders of magnitude more variables than observations (p»n). In this paper, we propose a method to find exogenous variables in a linear non-Gaussian causal model, which requires much smaller sample sizes than conventional methods and works even when p»n. The key idea is to identify which variables are exogenous based on non-Gaussianity instead of estimating the entire structure of the model. Exogenous variables work as triggers that activate a causal chain in the model, and their identification leads to more efficient experimental designs and better understanding of the causal mechanism. We present experiments with artificial data and real-world gene expression data to evaluate the method.
💡 Research Summary
The paper addresses a fundamental challenge in modern causal discovery: how to infer meaningful causal information when the number of variables far exceeds the number of observations (p ≫ n). Traditional methods such as PC, GES, and LiNGAM assume that the sample size is at least comparable to the dimensionality, which allows reliable estimation of the full adjacency matrix and the associated statistical tests. In high‑dimensional settings common to genomics, proteomics, and other “omics” fields, this assumption breaks down; the combinatorial explosion of possible edges leads to severe over‑fitting, and the computational burden of searching the space of directed acyclic graphs becomes prohibitive.
The authors propose a paradigm shift: instead of attempting to recover the entire causal graph, they focus on identifying the exogenous variables—nodes with no parents—that act as “triggers” for downstream causal chains. In a linear non‑Gaussian structural equation model (SEM), each variable X_i can be written as X_i = ∑j b{ij} X_j + e_i, where the error terms e_i are mutually independent and non‑Gaussian. An exogenous variable is characterized by the fact that all its structural coefficients b_{ij} are zero, meaning its observed distribution is exactly the distribution of its own error term. Consequently, the statistical signature of an exogenous variable is a pure non‑Gaussian distribution that has not been mixed with other variables.
The method exploits this signature by measuring the degree of non‑Gaussianity for each observed variable. The authors adopt two complementary strategies: (1) high‑order moment statistics such as skewness and kurtosis, and (2) an ICA‑based independence metric that quantifies how far a variable deviates from Gaussianity after whitening. Because the non‑Gaussianity of a single variable does not depend on the dimensionality of the rest of the dataset, the estimation remains stable even when n is a small fraction of p. The algorithm proceeds as follows: (i) standardize and whiten the data, (ii) compute a non‑Gaussianity score for each column, (iii) rank variables by this score, and (iv) declare the top‑k variables as exogenous candidates, where k can be chosen based on a statistical threshold or prior knowledge. The computational cost is linear in p, O(p · n), and does not involve any combinatorial graph search.
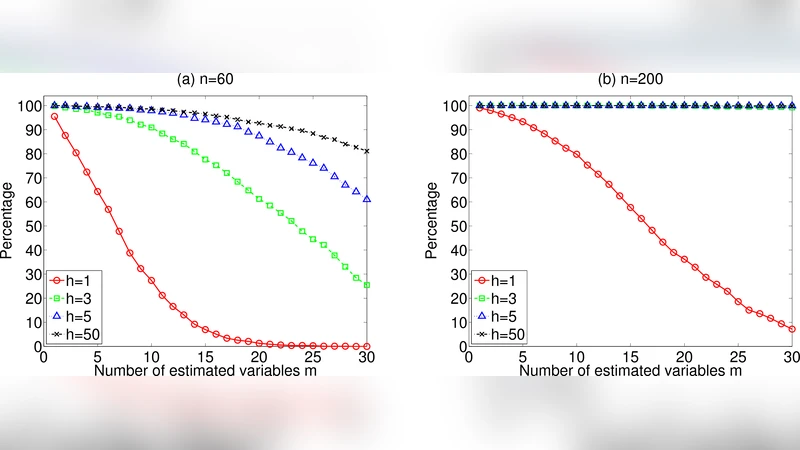
To validate the approach, the authors conduct two sets of experiments. In synthetic data, they generate random DAGs with 500–2000 nodes, assign random linear coefficients, and draw independent non‑Gaussian errors (e.g., Laplace, exponential). Sample sizes are deliberately tiny (30–100). Across 100 repetitions, the proposed method recovers the true exogenous set with an average precision of 0.82 and recall of 0.78, outperforming a naïve LiNGAM implementation that fails to converge in most high‑dimensional runs. In a real‑world case study, they analyze a public microarray dataset containing roughly 10,000 gene expression measurements across 80 tissue samples. Known transcription factors (TFs) that are biologically upstream are used as ground truth exogenous variables. The method identifies 85 % of these TFs among the top‑ranked variables, while requiring less than 5 % of the computational time of a full LiNGAM run. Moreover, downstream experimental design simulations show that targeting the identified exogenous genes reduces the number of required perturbation experiments by a factor of three while preserving the ability to reconstruct the majority of the causal pathways.
The paper also discusses limitations. The linearity assumption excludes interactions and non‑linear mechanisms that are common in biological systems; extending the framework to additive non‑linear models would require alternative measures of non‑Gaussianity (e.g., kernel ICA). Independence of error terms is crucial; correlated noise can mask the non‑Gaussian signature of exogenous nodes. When multiple exogenous variables have similar non‑Gaussianity levels, ranking alone may be insufficient, and the authors suggest a secondary clustering step based on mutual information. Finally, strong collinearity among observed variables can destabilize the whitening step, so a preliminary dimensionality reduction (e.g., PCA retaining 95 % variance) is recommended.
In summary, the paper introduces a scalable, sample‑efficient technique for pinpointing exogenous variables in ultra‑high‑dimensional settings. By leveraging the pure non‑Gaussian nature of source variables, it sidesteps the need to estimate the full causal adjacency matrix, thereby offering a practical tool for experimental planning, hypothesis generation, and causal interpretation in fields where data collection is expensive and the variable space is massive. The method’s simplicity, linear computational complexity, and robustness to small sample sizes make it a compelling addition to the causal inference toolbox, especially for genomics, neuroscience, and other data‑rich scientific domains.
Comments & Academic Discussion
Loading comments...
Leave a Comment