Applications of Littles Law to stochastic models of gene expression
The intrinsic stochasticity of gene expression can lead to large variations in protein levels across a population of cells. To explain this variability, different sources of mRNA fluctuations (‘Poisson’ and ‘Telegraph’ processes) have been proposed in stochastic models of gene expression. Both Poisson and Telegraph scenario models explain experimental observations of noise in protein levels in terms of ‘bursts’ of protein expression. Correspondingly, there is considerable interest in establishing relations between burst and steady-state protein distributions for general stochastic models of gene expression. In this work, we address this issue by considering a mapping between stochastic models of gene expression and problems of interest in queueing theory. By applying a general theorem from queueing theory, Little’s Law, we derive exact relations which connect burst and steady-state distribution means for models with arbitrary waiting-time distributions for arrival and degradation of mRNAs and proteins. The derived relations have implications for approaches to quantify the degree of transcriptional bursting and hence to discriminate between different sources of intrinsic noise in gene expression. To illustrate this, we consider a model for regulation of protein expression bursts by small RNAs. For a broad range of parameters, we derive analytical expressions (validated by stochastic simulations) for the mean protein levels as the levels of regulatory small RNAs are varied. The results obtained show that the degree of transcriptional bursting can, in principle, be determined from changes in mean steady-state protein levels for general stochastic models of gene expression.
💡 Research Summary
The paper addresses a fundamental problem in quantitative biology: how to relate the stochastic bursts of transcription to the steady‑state distribution of protein levels in cells. Two classic stochastic descriptions of transcription are considered – the Poisson model, where transcription events occur independently at a constant rate, and the Telegraph model, where a gene switches between an active “on” state and an inactive “off” state, producing bursts of mRNA when it is on. Both frameworks successfully capture the experimentally observed high variability (noise) in protein copy numbers, but they differ in how the burst size and frequency are defined.
The authors’ key insight is to map the biochemical processes of gene expression onto a queueing system. In this mapping, the synthesis of mRNA corresponds to the arrival of customers, the translation of mRNA into protein corresponds to the start of service, and the degradation of mRNA or protein corresponds to the completion of service. By doing so, the well‑known Little’s Law from queueing theory (L = λ W, where L is the average number of customers in the system, λ the arrival rate, and W the average time a customer spends in the system) can be applied to gene‑expression dynamics. Crucially, the authors prove a generalized version of Little’s Law that holds for arbitrary waiting‑time distributions, i.e., the inter‑arrival times of mRNA and the lifetimes of mRNA and protein need not be exponential.
Using this generalized theorem, they derive an exact relationship between the mean burst size (B), the mean inter‑burst interval (τ), and the mean steady‑state protein number ⟨P⟩:
⟨P⟩ = B · τ · ⟨Tₚ⟩
where ⟨Tₚ⟩ is the average protein lifetime. This formula is independent of the detailed shape of the underlying distributions, making it applicable to a wide class of stochastic gene‑expression models, including those with non‑Markovian waiting times.
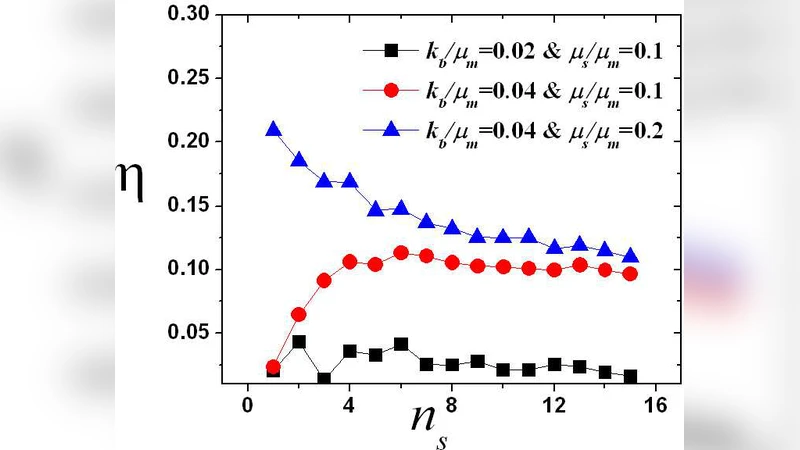
To demonstrate the practical utility of the result, the authors study a regulatory scenario in which small regulatory RNAs (sRNAs) bind to target mRNAs, forming complexes that either block translation or accelerate degradation. The model incorporates the sRNA concentration, binding and unbinding rates, and the degradation rate of the sRNA‑mRNA complex, each allowed to follow arbitrary distributions. Applying the derived Little’s‑Law relation, they obtain analytical expressions for the mean protein level as a function of sRNA concentration. These expressions predict a monotonic decrease of ⟨P⟩ with increasing sRNA, with a characteristic saturation at high sRNA levels. The analytical predictions are validated by extensive Gillespie stochastic simulations, showing excellent quantitative agreement.
An important implication of the work is that, because the mean protein level depends directly on the burst parameters, measuring only the average protein concentration under different regulatory conditions (e.g., varying sRNA levels) can be sufficient to infer the underlying transcriptional bursting strength. This provides a powerful, experimentally accessible method to distinguish between different sources of intrinsic noise (bursting versus Poissonian transcription) without requiring single‑molecule time‑course data.
The paper concludes by discussing the broader relevance of the approach. By extending Little’s Law to non‑exponential waiting times, the authors open the door to modeling more realistic cellular environments where degradation, processing, and transport steps have complex kinetics. The framework can be readily extended to multi‑gene networks, feedback loops, and other post‑transcriptional regulators. Moreover, the analytical tractability offered by the Little’s‑Law mapping makes it attractive for synthetic biology applications, where designers need predictable relationships between promoter architecture, burst characteristics, and output protein levels.
In summary, the study provides a rigorous, general, and experimentally relevant bridge between transcriptional burst statistics and steady‑state protein abundance, leveraging a classic theorem from queueing theory to advance our quantitative understanding of stochastic gene expression.
Comments & Academic Discussion
Loading comments...
Leave a Comment