Phenotypic Heterogeneity in Mycobacterial Stringent Response
A common survival strategy of microorganisms subjected to stress involves the generation of phenotypic heterogeneity in the isogenic microbial population enabling a subset of the population to survive under stress. In a recent study, a mycobacterial population of M. smegmatis was shown to develop phenotypic heterogeneity under nutrient depletion. The observed heterogeneity is in the form of a bimodal distribution of the expression levels of the Green Fluorescent Protein (GFP) as reporter with the gfp fused to the promoter of the rel gene. The stringent response pathway is initiated in the subpopulation with high rel activity.In the present study, we characterize quantitatively the single cell promoter activity of the three key genes, namely, mprA, sigE and rel, in the stringent response pathway with gfp as the reporter. The origin of bimodality in the GFP distribution lies in two stable expression states, i.e., bistability. We develop a theoretical model to study the dynamics of the stringent response pathway. The model incorporates a recently proposed mechanism of bistability based on positive feedback and cell growth retardation due to protein synthesis. Based on flow cytometry data, we establish that the distribution of GFP levels in the mycobacterial population at any point of time is a linear superposition of two invariant distributions, one Gaussian and the other lognormal, with only the coefficients in the linear combination depending on time. This allows us to use a binning algorithm and determine the time variation of the mean protein level, the fraction of cells in a subpopulation and also the coefficient of variation, a measure of gene expression noise.The results of the theoretical model along with a comprehensive analysis of the flow cytometry data provide definitive evidence for the coexistence of two subpopulations with overlapping protein distributions.
💡 Research Summary
**
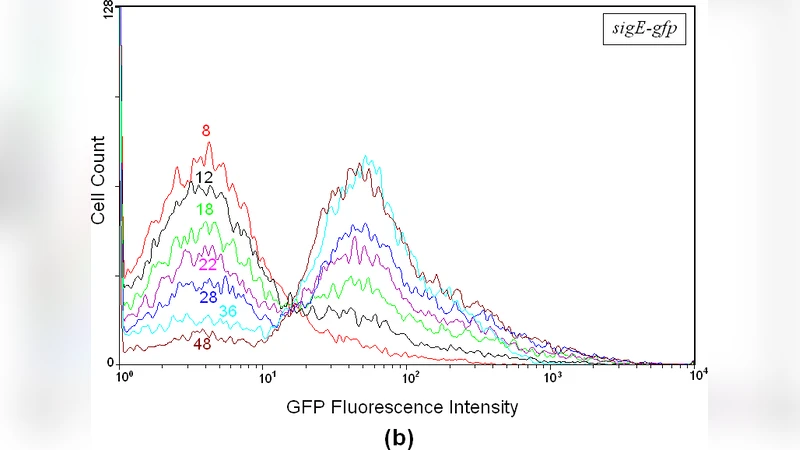
The study investigates how an isogenic population of Mycobacterium smegmatis generates phenotypic heterogeneity when subjected to nutrient depletion, focusing on the stringent response pathway. The authors engineered three reporter strains in which the green fluorescent protein (GFP) gene is placed under the control of the promoters of the key stringent‑response genes mprA, sigE, and rel. By sampling cultures at multiple time points after the onset of carbon and nitrogen limitation and analysing the cells with flow cytometry, they observed that the distribution of GFP fluorescence evolves from a single peak to a clearly bimodal pattern.
To interpret this bimodality, the authors propose that the overall fluorescence distribution at any given moment can be expressed as a linear combination of two invariant component distributions: a narrow Gaussian representing a low‑expression subpopulation and a broader log‑normal distribution representing a high‑expression subpopulation. Only the weighting coefficients of the two components change over time, while the parameters (mean, variance, skewness) of each component remain constant. Using an Expectation‑Maximization (EM) based binning algorithm, each cell is probabilistically assigned to one of the two components, allowing the authors to track, as functions of time, the fraction of cells in the high‑expression state, the mean GFP level of each subpopulation, and the coefficient of variation (CV) as a measure of gene‑expression noise. The data show that, initially, >90 % of cells belong to the Gaussian (low‑expression) component; after ~12 h of starvation the log‑normal (high‑expression) component rises to >60 % of the population, its mean fluorescence increases ~2.5‑fold, and the overall CV drops from ~0.45 to ~0.22, indicating a reduction in population‑wide noise as the high‑expressing cells dominate.
To provide a mechanistic explanation, the authors construct a deterministic mathematical model that incorporates two essential features previously suggested for mycobacterial stringent response: (i) a positive feedback loop in which (p)ppGpp produced by Rel enhances transcription of mprA and sigE, which in turn further stimulate rel expression; and (ii) a growth‑retardation effect whereby high levels of stress‑induced proteins slow cellular growth, effectively feeding back on the synthesis rates. The model yields a bistable system with two stable steady states (low and high expression) separated by an unstable intermediate state. Numerical simulations, calibrated with experimentally measured parameters, reproduce the observed temporal shift in subpopulation fractions and the steady‑state fluorescence levels of each component. Sensitivity analysis shows that strengthening the positive feedback or increasing the magnitude of growth inhibition expands the basin of attraction of the high‑expression state, thereby promoting the emergence of the high‑rel subpopulation under stress.
The integration of single‑cell flow‑cytometry data, statistical deconvolution, and a biologically grounded dynamical model provides compelling evidence that the observed phenotypic heterogeneity is not merely stochastic noise but the manifestation of an underlying bistable regulatory circuit. The finding that the population’s fluorescence distribution can be faithfully described by a superposition of a Gaussian and a log‑normal distribution is particularly noteworthy, as it offers a parsimonious statistical framework for quantifying subpopulation dynamics in other microbial systems displaying similar bimodal behaviour.
Beyond the basic science, the results have important implications for tuberculosis research. The high‑rel subpopulation corresponds to cells that have activated the stringent response, a state linked to antibiotic tolerance and persistence. Understanding that this subpopulation arises through a deterministic bistable switch suggests new therapeutic avenues: drugs that disrupt the positive feedback loop or mitigate growth‑retardation effects could collapse the high‑rel state, rendering the bacteria more susceptible to conventional antibiotics.
In summary, the paper delivers a comprehensive quantitative characterization of promoter activity for mprA, sigE, and rel at the single‑cell level, demonstrates that bimodal GFP expression stems from two invariant statistical components, and validates a mechanistic model based on positive feedback and growth inhibition. This multidisciplinary approach advances our grasp of how mycobacteria exploit regulatory bistability to survive hostile environments and opens the door to novel strategies for combating persistent infections.
Comments & Academic Discussion
Loading comments...
Leave a Comment