Spatial clustering of array CGH features in combination with hierarchical multiple testing
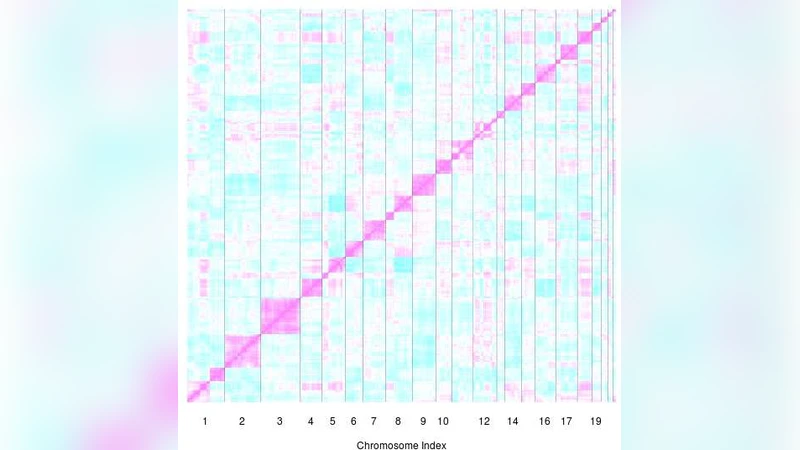
We propose a new approach for clustering DNA features using array CGH data from multiple tumor samples. We distinguish data-collapsing: joining contiguous DNA clones or probes with extremely similar data into regions, from clustering: joining contiguous, correlated regions based on a maximum likelihood principle. The model-based clustering algorithm accounts for the apparent spatial patterns in the data. We evaluate the randomness of the clustering result by a cluster stability score in combination with cross-validation. Moreover, we argue that the clustering really captures spatial genomic dependency by showing that coincidental clustering of independent regions is very unlikely. Using the region and cluster information, we combine testing of these for association with a clinical variable in an hierarchical multiple testing approach. This allows for interpreting the significance of both regions and clusters while controlling the Family-Wise Error Rate simultaneously. We prove that in the context of permutation tests and permutation-invariant clusters it is allowed to perform clustering and testing on the same data set. Our procedures are illustrated on two cancer data sets.
💡 Research Summary
The paper introduces a comprehensive statistical framework for analyzing spatial patterns in array CGH data obtained from multiple tumor samples. The authors first distinguish two preprocessing steps: data‑collapsing and clustering. In the data‑collapsing stage, contiguous probes or clones that exhibit nearly identical copy‑number profiles are merged into single “regions.” This reduces dimensionality, mitigates noise, and creates a set of regions that each represent a homogeneous genomic segment. The second stage applies a model‑based clustering algorithm that groups adjacent regions into larger “clusters” according to a maximum‑likelihood principle. Each region’s copy‑number measurements are assumed to follow a multivariate normal distribution, and the covariance structure is parameterized to reflect spatial proximity, often using distance‑based weighting. By maximizing the overall log‑likelihood, the algorithm identifies clusters that capture genuine spatial dependency rather than random co‑occurrence.
To assess whether the observed clustering could arise by chance, the authors develop a cluster stability score. They repeatedly resample the data (via bootstrap or cross‑validation), re‑run the clustering, and compare the resulting partitions to the original using similarity metrics such as the Jaccard index. High average similarity indicates that the cluster configuration is reproducible and not an artifact of sampling variability. Additionally, they analytically compute the probability that independent regions would be grouped together by random chance, demonstrating that the observed clustering is statistically unlikely under a null model of independence.
The central methodological contribution is a hierarchical multiple‑testing procedure that integrates region‑level and cluster‑level inference while controlling the Family‑Wise Error Rate (FWER). The authors perform permutation tests for association between each region’s copy‑number profile and a clinical outcome (e.g., survival, treatment response). Because the clustering is permutation‑invariant—clusters remain the same under any permutation of sample labels—they can safely recompute clusters for each permuted dataset. For each permutation, the smallest p‑value among regions within a cluster becomes the cluster‑level p‑value. Applying a Westfall‑Young step‑down adjustment across both levels yields a single set of adjusted p‑values that simultaneously control the FWER for all region and cluster hypotheses. This theoretical result guarantees that clustering and testing on the same dataset do not inflate type‑I error, provided the permutation framework is respected.
The authors illustrate the approach on two cancer data sets, one breast‑cancer and one lung‑cancer cohort. In both cases, the method identifies fewer but more biologically coherent regions than traditional probe‑wise testing. Importantly, the regions cluster into genomic segments that correspond to known hotspots of copy‑number amplification (e.g., 8q24) or deletion. These clusters show strong, statistically significant associations with patient outcomes, often surpassing the predictive power of any single region. The stability scores for the clusters are high, confirming that the spatial structure is robust across resampling.
In summary, the paper delivers (1) a clear separation of data‑collapsing and spatial clustering, (2) quantitative tools for evaluating cluster reproducibility and randomness, and (3) a rigorous hierarchical permutation‑based multiple‑testing scheme that controls FWER while leveraging spatial dependency. The methodology is not limited to array CGH; it can be extended to other high‑density genomic platforms such as SNP arrays or methylation arrays, offering a powerful way to uncover clinically relevant genomic patterns that respect the underlying spatial organization of the genome.
Comments & Academic Discussion
Loading comments...
Leave a Comment