Inference of plasmid copy number mean and noise from single cell gene expression data
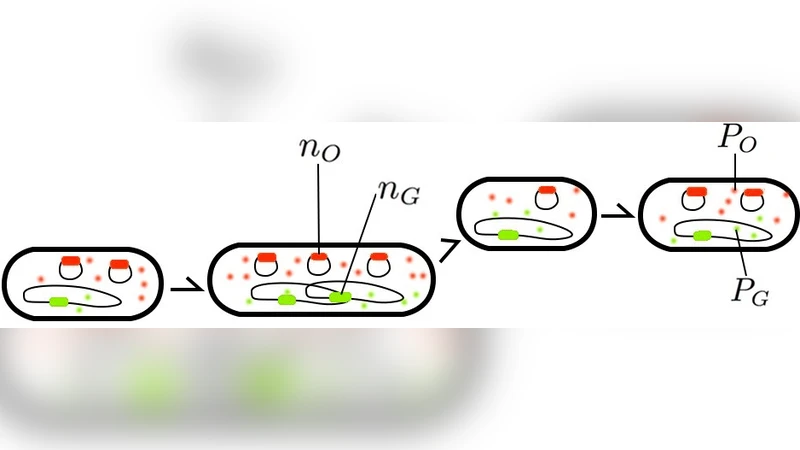
Plasmids are extra-chromosomal DNA molecules which code for their own replication. We previously reported a setup using genes coding for fluorescent proteins of two colors that allowed us, using a simple model, to extract the plasmid copy number noise in a monoclonal population of bacteria [J. Wong Ng et al., Phys. Rev. E, 81, 011909 (2010)]. Here we present a detailed calculation relating this noise to the measured levels of fluorescence, taking into account all sources of fluorescence fluctuations: the fluctuation of gene expression as in the simple model, but also the growth and division of bacteria, the non-uniform distribution of their ages, the random partition of proteins at divisions and the replication and partition of plasmids and chromosome. We show how using the chromosome as a reference helps extracting the plasmid copy number noise in a self-consistent manner.
💡 Research Summary
Plasmids are widely used as vectors in bacterial genetics, yet quantitative knowledge of their copy‑number distribution within a population remains difficult to obtain. Traditional methods such as quantitative PCR or electron microscopy are labor‑intensive and provide only bulk‑average information. In this paper the authors present a rigorous statistical framework that extracts both the mean plasmid copy number and its cell‑to‑cell variability (noise) from single‑cell fluorescence measurements, while explicitly accounting for all major sources of variability in a growing bacterial culture.
The experimental design relies on two fluorescent reporters driven by identical promoters and ribosome‑binding sites. One reporter (e.g., GFP) is encoded on the plasmid of interest, the other (e.g., mCherry) is integrated into the chromosome at a neutral locus. Because the transcription‑translation machinery is shared, any systematic difference between the two fluorescence signals can be attributed to differences in gene dosage, i.e., the number of plasmid copies versus the single chromosomal copy.
To translate raw fluorescence into copy‑number statistics the authors construct a comprehensive stochastic model. The model includes:
- Intrinsic and extrinsic noise of gene expression, modeled as a combination of Poisson (transcription) and Gaussian (translation/measurement) fluctuations.
- Cell growth and binary division, where proteins are partitioned randomly between daughter cells according to a Bernoulli process (probability ½).
- A non‑uniform age distribution of cells in an exponentially growing culture, described by an exponential density λ(a)=2e⁻²ᵃ for normalized age a∈
Comments & Academic Discussion
Loading comments...
Leave a Comment