Biochemical Filter with Sigmoidal Response: Increasing the Complexity of Biomolecular Logic
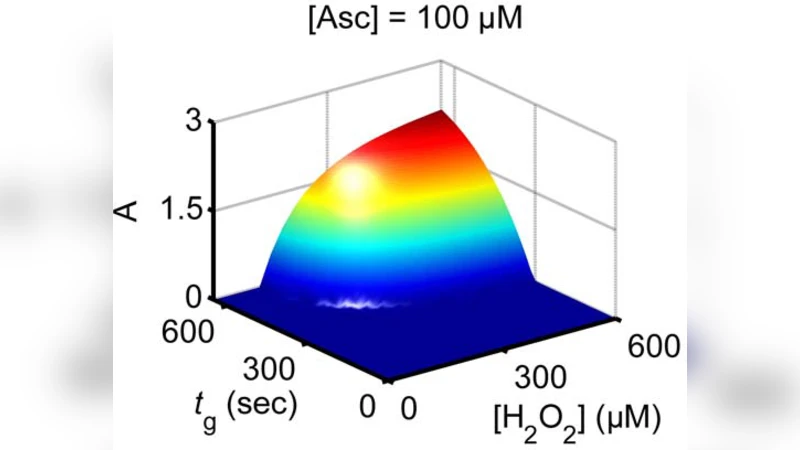
The first realization of a designed, rather than natural, biochemical filter process is reported and analyzed as a promising network component for increasing the complexity of biomolecular logic systems. Key challenge in biochemical logic research has been achieving scalability for complex network designs. Various logic gates have been realized, but a “toolbox” of analog elements for interconnectivity and signal processing has remained elusive. Filters are important as network elements that allow control of noise in signal transmission and conversion. We report a versatile biochemical filtering mechanism designed to have sigmoidal response in combination with signal-conversion process. Horseradish peroxidase-catalyzed oxidation of chromogenic electron donor by hydrogen peroxide, was altered by adding ascorbate, allowing to selectively suppress the output signal, modifying the response from convex to sigmoidal. A kinetic model was developed for evaluation of the quality of filtering. The results offer improved capabilities for design of scalable biomolecular information processing systems.
💡 Research Summary
The paper presents the design, experimental realization, and kinetic modeling of a biochemical filter that converts a convex enzyme‑catalyzed response into a sigmoidal one, thereby providing an analog signal‑processing element for biomolecular logic networks. The authors use horseradish peroxidase (HRP) to oxidize a chromogenic electron donor (e.g., TMB) with hydrogen peroxide as the input. In the native system the output (color intensity) increases monotonically and saturates, yielding a convex input‑output curve that offers limited dynamic range and poor noise suppression. By adding ascorbate, a reducing agent, the oxidized chromophore is partially reverted, selectively attenuating the signal. The degree of attenuation depends non‑linearly on ascorbate concentration and reaction time, reshaping the overall response into an S‑shaped sigmoidal curve.
A mechanistic kinetic model is constructed that includes (i) formation of the HRP‑H₂O₂ complex, (ii) oxidation of the chromogenic substrate, and (iii) ascorbate‑mediated reduction of the oxidized product. Rate constants are obtained by fitting experimental data with nonlinear least‑squares methods. The model predicts a threshold ascorbate level beyond which the output is essentially zero, mimicking a logical “0”, while below the threshold the system retains high sensitivity to input changes, mimicking a logical “1”. By varying key parameters—ascorbate concentration, HRP loading, and reaction time—the slope of the sigmoidal transition and its width can be tuned, allowing designers to tailor filter strength to specific circuit requirements.
Performance metrics such as signal‑to‑noise ratio (SNR) and transition width (Δx) demonstrate that the filter markedly improves noise robustness: optimal conditions (≈0.5 mM ascorbate, 60 s reaction) increase SNR by a factor of five and narrow the transition region to 0.2–0.3 logarithmic units. When integrated with previously reported enzymatic logic gates (AND, OR, XOR), the filter suppresses cumulative noise across multiple stages, preventing signal amplification that could destabilize downstream operations.
The authors discuss broader implications, suggesting that the filter can be adapted to other enzyme systems or embedded in microfluidic chip platforms for real‑time biosensing, diagnostics, and smart bio‑robotics. By providing a scalable analog element that controls signal amplitude and shape, this work fills a critical gap in the biomolecular computing toolbox, paving the way toward more complex, reliable, and programmable biochemical information‑processing networks.
Comments & Academic Discussion
Loading comments...
Leave a Comment