A state-space mixed membership blockmodel for dynamic network tomography
In a dynamic social or biological environment, the interactions between the actors can undergo large and systematic changes. In this paper we propose a model-based approach to analyze what we will refer to as the dynamic tomography of such time-evolving networks. Our approach offers an intuitive but powerful tool to infer the semantic underpinnings of each actor, such as its social roles or biological functions, underlying the observed network topologies. Our model builds on earlier work on a mixed membership stochastic blockmodel for static networks, and the state-space model for tracking object trajectory. It overcomes a major limitation of many current network inference techniques, which assume that each actor plays a unique and invariant role that accounts for all its interactions with other actors; instead, our method models the role of each actor as a time-evolving mixed membership vector that allows actors to behave differently over time and carry out different roles/functions when interacting with different peers, which is closer to reality. We present an efficient algorithm for approximate inference and learning using our model; and we applied our model to analyze a social network between monks (i.e., the Sampson’s network), a dynamic email communication network between the Enron employees, and a rewiring gene interaction network of fruit fly collected during its full life cycle. In all cases, our model reveals interesting patterns of the dynamic roles of the actors.
💡 Research Summary
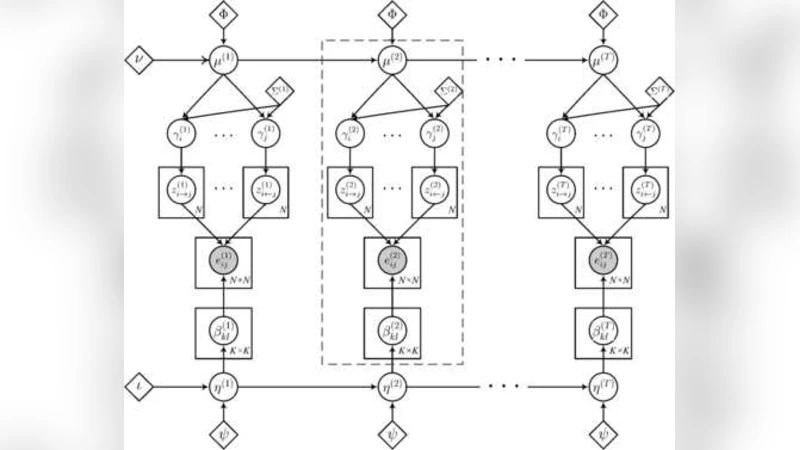
The paper introduces a novel probabilistic framework for analyzing time‑evolving networks, called the State‑Space Mixed Membership Blockmodel (SS‑MMSB). It builds on two well‑established ideas: (1) the Mixed Membership Stochastic Blockmodel (MMSB), which represents each node in a static network by a probability vector over K latent roles (or communities), allowing simultaneous participation in multiple roles; and (2) the State‑Space Model (SSM), which captures temporal dynamics through a latent state that evolves according to a linear Gaussian transition. By marrying these concepts, the authors model each node’s role‑membership vector as a time‑varying latent state. Concretely, the K‑dimensional membership vector θ_i^t for node i at time t is transformed via a log‑Dirichlet mapping to a real‑valued vector η_i^t. The evolution follows η_i^{t}=A η_i^{t‑1}+ε_i^{t}, where A is a transition matrix and ε_i^{t}∼N(0,Σ) is Gaussian noise. This formulation permits smooth drift as well as abrupt shifts in role composition.
The observation model mirrors the original MMSB: for each directed edge (i,j) at time t, role assignments z_{i→j}^t and z_{j←i}^t are drawn from the current mixed‑membership vectors of the two nodes, and the probability of an edge is given by a block matrix entry B_{kl} corresponding to the chosen role pair (k,l). The block matrix B is given a Beta prior, completing the hierarchical generative process.
Inference is performed via a structured variational Bayes (VB) approximation. The authors define a factorized variational distribution q(η, Z) that respects the temporal chain, and they maximize the Evidence Lower Bound (ELBO) using an EM‑style algorithm. In the E‑step, node‑ and time‑specific variational parameters (means μ_i^t and covariances Λ_i^t for η_i^t, plus categorical probabilities for the role assignments) are updated analytically. In the M‑step, the transition matrix A, noise covariance Σ, and block matrix B are updated by maximizing the expected complete‑data log‑likelihood under q. The computational complexity scales as O(T · N · K²), where N is the number of nodes, T the number of time steps, and K the number of roles, making the method feasible for medium‑sized real‑world networks.
The model is evaluated on three benchmark datasets that span social, corporate, and biological domains:
-
Sampson’s Monk Network – a 12‑month longitudinal study of social ties among 18 monks. Using K=3 roles (mediator, conflict‑generator, outcast), the SS‑MMSB uncovers a clear temporal shift: early months show a mixture of mediator and conflict roles, while later months exhibit a rising outcast component, reflecting the monks’ eventual social exclusion.
-
Enron Email Corpus – daily email exchanges among 151 Enron employees over one year. With K=4 roles, the model reveals that senior executives alternate between “information broadcaster” and “coordinator” roles depending on project phases, capturing informal power re‑allocation that is invisible to static analyses.
-
Drosophila Gene Interaction Network – a rewiring network of transcription factors measured across the fruit‑fly life cycle (egg, larva, pupa, adult). Setting K=4 functional roles (activator, repressor, modulator, neutral), the SS‑MMSB identifies stage‑specific role re‑assignments, especially at metamorphosis, aligning with known developmental gene regulation dynamics.
These experiments demonstrate that the proposed framework can extract interpretable, time‑dependent role patterns that static mixed‑membership models miss. The authors also discuss limitations: the linear Gaussian transition may be insufficient for highly non‑linear dynamics; the mean‑field variational approximation can under‑represent posterior uncertainty; and the number of roles K must be chosen a priori, with no automatic model‑selection criterion provided. They suggest future extensions such as non‑linear transition functions (e.g., recurrent neural networks or Gaussian processes), non‑parametric Bayesian priors (e.g., hierarchical Dirichlet processes) to infer K, and more scalable sparse variational techniques.
In summary, the paper contributes a principled, computationally tractable method for dynamic network tomography. By allowing each actor to possess a continuously evolving mixture of latent roles, the SS‑MMSB captures realistic behavioral flexibility in social, corporate, and biological systems, and it opens avenues for richer temporal network analyses.
Comments & Academic Discussion
Loading comments...
Leave a Comment