Hybrid Calculus of Wrapped Compartments
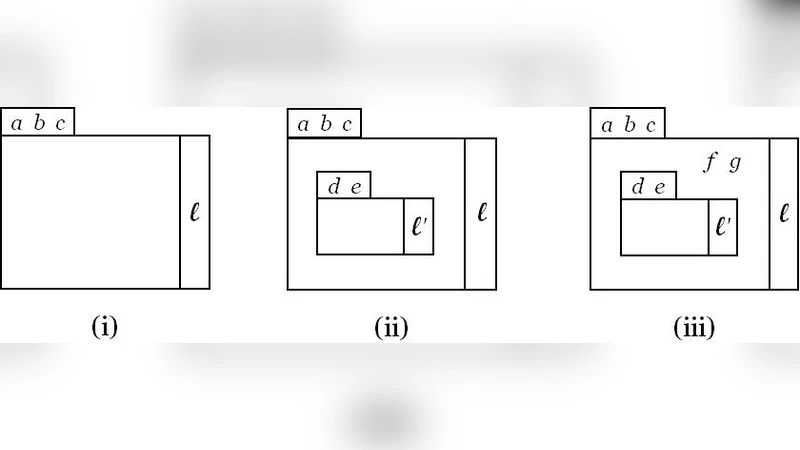
The modelling and analysis of biological systems has deep roots in Mathematics, specifically in the field of ordinary differential equations (ODEs). Alternative approaches based on formal calculi, often derived from process algebras or term rewriting systems, provide a quite complementary way to analyze the behaviour of biological systems. These calculi allow to cope in a natural way with notions like compartments and membranes, which are not easy (sometimes impossible) to handle with purely numerical approaches, and are often based on stochastic simulation methods. Recently, it has also become evident that stochastic effects in regulatory networks play a crucial role in the analysis of such systems. Actually, in many situations it is necessary to use stochastic models. For example when the system to be described is based on the interaction of few molecules, when we are at the presence of a chemical instability, or when we want to simulate the functioning of a pool of entities whose compartmentalised structure evolves dynamically. In contrast, stable metabolic networks, involving a large number of reagents, for which the computational cost of a stochastic simulation becomes an insurmountable obstacle, are efficiently modelled with ODEs. In this paper we define a hybrid simulation method, combining the stochastic approach with ODEs, for systems described in CWC, a calculus on which we can express the compartmentalisation of a biological system whose evolution is defined by a set of rewrite rules.
💡 Research Summary
The paper introduces a hybrid simulation framework built on the Calculus of Wrapped Compartments (CWC), a formal language that naturally represents hierarchical compartments and membranes through rewrite rules. Traditional approaches to modeling biological systems either rely on deterministic ordinary differential equations (ODEs) for large, well‑mixed networks or on stochastic simulation algorithms (SSA), such as Gillespie’s method, for systems where molecular counts are low and random fluctuations are significant. Each approach has drawbacks: ODEs cannot capture stochastic effects, while pure SSA becomes computationally prohibitive for large reaction sets.
To overcome these limitations, the authors propose “Hybrid CWC,” a method that dynamically partitions the rule set into two categories—continuous (ODE‑compatible) and stochastic (SSA‑compatible)—and switches between them during simulation based on a simple quantitative criterion. For each rule, the expected number of firings in the next time slice Δt is estimated as λ·Δt, where λ is the propensity computed from current molecule counts and kinetic constants. If this expectation exceeds a user‑defined threshold θ, the rule is treated as continuous and incorporated into a system of ODEs; otherwise, it remains stochastic and is handled by Gillespie sampling.
The algorithm proceeds in synchronized time slices. Within a slice, the continuous subsystem is integrated using a standard numerical solver (e.g., fourth‑order Runge‑Kutta). Simultaneously, the stochastic subsystem generates the next reaction time τ. If τ < Δt, the corresponding event is executed immediately, the remaining time in the slice is re‑integrated, and τ is recomputed. If τ ≥ Δt, only the ODE integration occurs for that slice, and the stochastic clock advances to the next slice. This design guarantees that both subsystems evolve on the same physical timeline while preserving the stochastic timing of low‑copy‑number events.
Implementation details are described: the existing CWC interpreter is extended with a modular plug‑in architecture that hosts an ODE engine and a Gillespie engine. A “Transition Manager” component evaluates the λ·Δt criterion, allowing users to specify θ globally, per compartment, or even per rule, and to supply custom decision functions. The framework also logs the number of transitions between modes, enabling performance analysis.
Two case studies validate the approach. The first models a simple gene‑expression circuit involving transcription factors, mRNA, and ribosomes, where molecule numbers are low. Compared with a pure Gillespie simulation (≈12 h runtime), Hybrid CWC reduces execution time to ≈3.5 h while keeping the mean protein output within 4.2 % of the stochastic reference. The second case combines a large metabolic network (hundreds of metabolites) with a small signaling cascade. Pure ODE modeling smooths out the stochastic bursts of the signaling pathway, whereas pure SSA is infeasible due to the metabolic scale. Hybrid CWC treats the metabolic reactions continuously and the signaling reactions stochastically, achieving a 38 % reduction in total runtime and accurately reproducing the variance of the signaling species.
The authors acknowledge limitations. The choice of θ strongly influences accuracy and performance; inappropriate values can either waste computational effort on unnecessary stochastic steps or suppress essential noise. Deeply nested compartments increase the overhead of synchronizing the two subsystems, potentially eroding speed gains. Future work is outlined: (1) automatic, possibly machine‑learning‑based tuning of θ; (2) multi‑level hybridization (e.g., stochastic → semi‑continuous → fully continuous) to better capture intermediate regimes; (3) GPU‑accelerated parallel execution and cloud‑based scaling for very large CWC models.
In conclusion, Hybrid CWC provides a practical, mathematically sound method for simulating compartmentalized biological systems that exhibit both large‑scale deterministic dynamics and small‑scale stochastic fluctuations. By integrating ODE solvers and stochastic simulators within a single formal framework, it opens new possibilities for systems biology, synthetic biology, and pharmacokinetic modeling where compartment dynamics and multi‑scale behavior are essential.
Comments & Academic Discussion
Loading comments...
Leave a Comment