Population Structure and Cryptic Relatedness in Genetic Association Studies
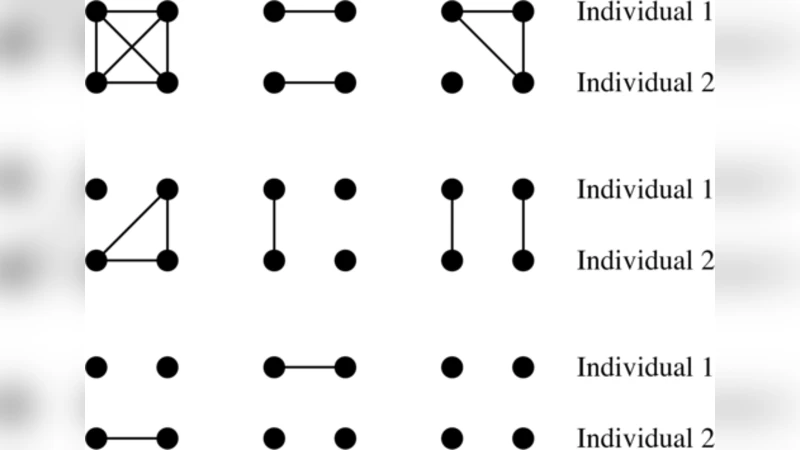
We review the problem of confounding in genetic association studies, which arises principally because of population structure and cryptic relatedness. Many treatments of the problem consider only a simple ``island’’ model of population structure. We take a broader approach, which views population structure and cryptic relatedness as different aspects of a single confounder: the unobserved pedigree defining the (often distant) relationships among the study subjects. Kinship is therefore a central concept, and we review methods of defining and estimating kinship coefficients, both pedigree-based and marker-based. In this unified framework we review solutions to the problem of population structure, including family-based study designs, genomic control, structured association, regression control, principal components adjustment and linear mixed models. The last solution makes the most explicit use of the kinships among the study subjects, and has an established role in the analysis of animal and plant breeding studies. Recent computational developments mean that analyses of human genetic association data are beginning to benefit from its powerful tests for association, which protect against population structure and cryptic kinship, as well as intermediate levels of confounding by the pedigree.
💡 Research Summary
This paper addresses a central source of bias in genetic association studies: confounding caused by population structure and cryptic relatedness. Rather than treating these phenomena as separate problems, the authors propose a unified perspective in which both are manifestations of a single, unobserved pedigree that defines the (often distant) relationships among study participants. The pedigree, or kinship, becomes the core concept around which the entire review is organized.
First, the authors clarify the definition of the kinship coefficient and discuss two major families of estimation methods. Pedigree‑based approaches use known family trees to compute exact coefficients, but they are limited when genealogical information is incomplete or unavailable. Marker‑based methods, which infer kinship directly from genome‑wide SNP data, are described in detail: method‑of‑moments estimators, maximum‑likelihood formulations, and Bayesian frameworks. These approaches can capture distant, otherwise invisible relationships and thus are essential for detecting cryptic relatedness in large, ostensibly unrelated cohorts.
The paper then surveys a broad spectrum of strategies that have been proposed to correct for confounding. Family‑based designs (e.g., trio or sib‑pair studies) eliminate bias at the source by leveraging explicit pedigree information, yet they are costly and often underpowered for complex traits. Genomic control rescales test statistics using a global inflation factor (λ), providing a simple but coarse correction that fails to address local structure. Structured association methods (e.g., STRUCTURE, ADMIXTURE) and regression‑based adjustments incorporate inferred ancestry clusters or covariates, but they depend critically on the correct specification of the number of clusters and may over‑ or under‑correct. Principal components analysis (PCA) extracts the leading axes of genetic variation and uses them as covariates; while computationally efficient, PCA does not fully account for subtle kinship‑induced correlations.
The most comprehensive solution presented is the linear mixed model (LMM). In an LMM, the phenotype is modeled as a sum of fixed effects (including the SNP of interest) and a random effect whose covariance structure is proportional to the kinship matrix K. This formulation simultaneously controls for population structure (captured by the eigenvectors of K) and cryptic relatedness (captured by the off‑diagonal elements of K). The authors review several algorithmic advances that have made LMMs tractable for modern GWAS: EMMA, FastLMM, GEMMA, REGENIE, and related methods that exploit spectral decomposition, low‑rank approximations, and efficient gradient calculations. These tools reduce computational complexity from O(N³) to O(N²) or even O(N) for very large sample sizes, enabling the analysis of hundreds of thousands of individuals and millions of markers.
Beyond algorithmic speed, the paper emphasizes the statistical advantages of LMMs. By explicitly modeling the genetic relatedness matrix, LMMs provide unbiased estimates of SNP effects even in the presence of complex, hierarchical population structure. They also improve power relative to simpler methods because the random effect absorbs background polygenic variation, reducing residual variance. The authors illustrate these points with examples from animal and plant breeding, where LMMs have long been standard, and show how recent implementations are now being applied to human data with comparable success.
Practical guidance is offered for researchers embarking on GWAS. The authors recommend rigorous quality control (QC) of genotype data, LD pruning before kinship estimation, and visual inspection of the estimated kinship matrix (e.g., heatmaps) to detect unexpected close relationships. They suggest a stepwise workflow: start with PCA or genomic control for a quick sanity check, then compute a marker‑based kinship matrix and fit an LMM for the final association tests. Model diagnostics—such as QQ plots, inspection of the λ inflation factor after LMM fitting, and cross‑validation of significant hits—are highlighted as essential to ensure that confounding has been adequately addressed.
In conclusion, the review reframes population structure and cryptic relatedness as two sides of the same pedigree‑based confounder, positions kinship as the unifying metric, and demonstrates that linear mixed models, empowered by recent computational breakthroughs, constitute the most robust and versatile framework for modern genetic association studies. By integrating pedigree information—whether explicit or inferred from markers—researchers can achieve reliable association signals while protecting against both overt and subtle sources of bias.
Comments & Academic Discussion
Loading comments...
Leave a Comment