Unwinding dynamics of double-stranded polymers
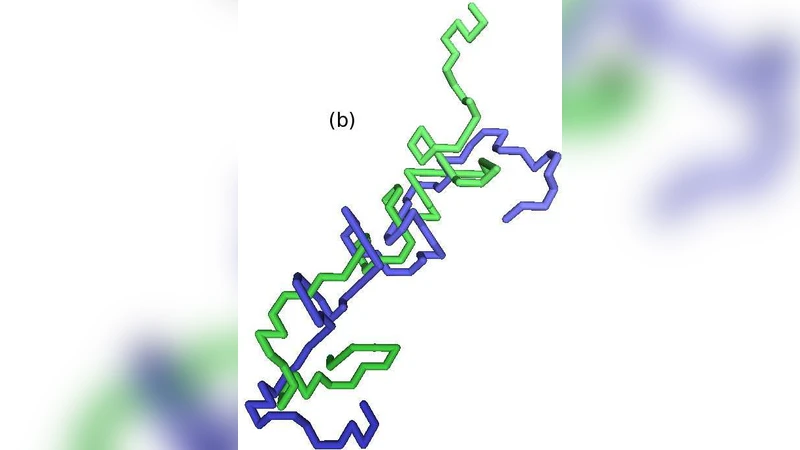
We consider the unwinding of two lattice polymer strands of length N that are initially wound around each other in a double-helical conformation and evolve through Rouse dynamics. The problem relates to quickly bringing a double-stranded polymer well above its melting temperature, i.e., the binding interactions between the strands are neglected, and the strands separate from each other as it is entropically favorable for them to do so. The strands unwind by rotating around each other until they separate. We find that the process proceeds from the ends inward; intermediate conformations can be characterized by a tightly wound inner part, from which loose strands are sticking out, with length lt^0.39. The total time needed for the two strands to unwind scales as a power of N as tuN^(2.57+-0.03). We present a theoretical argument, which suggests that during this unwinding process, these loose strands are far out of equilibrium.
💡 Research Summary
The paper investigates the dynamics of unwinding two lattice polymer strands that are initially intertwined in a double‑helical configuration and then evolve under pure Rouse dynamics, i.e., without any attractive interactions between the strands. This situation mimics the rapid heating of a double‑stranded polymer above its melting temperature, where the binding forces become negligible and entropy drives the strands apart. The authors use Monte‑Carlo simulations of self‑avoiding walks on a three‑dimensional cubic lattice for various chain lengths N (from 64 up to 1024 monomers). Each monomer follows overdamped Langevin dynamics with stochastic thermal kicks and viscous drag, reproducing the classic Rouse model.
The key observations are:
-
End‑initiated unwinding – The separation starts at the free ends of the two chains and proceeds inward. The interior remains a tightly wound “core” while the outer portions become loose, dangling strands.
-
Growth law for the loose segment – The average length ℓ(t) of the detached, loosely coiled part follows a sub‑diffusive power law ℓ ∝ t^0.39. This exponent is smaller than the ½ expected for simple diffusion, indicating that the unwinding is slowed by rotational constraints and the torque transmitted through the remaining helical core.
-
Overall unwinding time scaling – The total time τ_u required for the two polymers to become completely separated scales with the polymer length as τ_u ∝ N^α with α = 2.57 ± 0.03. This exponent exceeds the N^2 scaling of the ordinary Rouse relaxation time, reflecting the additional time cost associated with rotating the tightly wound core.
To rationalize these findings, the authors propose a simple analytical picture. At any moment the system consists of a compact helical core of length L_c(t) and two free tails of length ℓ(t) attached at the ends. The tails behave almost like free Rouse chains, but the torque τ_rot that must be supplied to rotate the core is inversely proportional to the tail length, τ_rot ∝ 1/ℓ. Consequently, as the tails grow, the rotational resistance diminishes, allowing the core to unwind faster. By assuming that the core shrinks when the torque exceeds a threshold, one obtains a differential equation dℓ/dt ∝ ℓ^–β, whose solution gives ℓ ∝ t^{1/(1+β)}. Matching the observed exponent 0.39 yields β ≈ 1.56. Substituting this β into the expression for the total unwinding time gives τ_u ∝ N^{2+β/(1+β)} ≈ N^{2.57}, in excellent agreement with the simulation data.
An important implication of the analysis is that the unwinding process is intrinsically out‑of‑equilibrium. While the detached tails quickly acquire the statistics of a free polymer, the coupling to the rotating core maintains a persistent torque, preventing the whole system from reaching a global equilibrium configuration. This challenges the common assumption in polymer physics that local segments equilibrate instantaneously during large‑scale conformational changes.
The authors discuss experimental relevance. Rapid temperature jumps in DNA or RNA experiments, observed with single‑molecule fluorescence or magnetic tweezers, could be used to test the predicted ℓ ∝ t^0.39 law and the N^2.57 scaling of the total denaturation time. Moreover, the present model neglects residual hydrogen bonding or electrostatic interactions that would be present in real biomolecules; incorporating such forces would likely increase the effective exponent, offering a systematic way to quantify the strength of residual binding from kinetic measurements.
In summary, the paper provides a clear computational and theoretical framework for the unwinding of double‑helical polymers in the high‑temperature, non‑binding limit. It demonstrates that unwinding proceeds from the ends inward, that the length of the loose tails grows sub‑diffusively with time, and that the total unwinding time scales as N^2.57. The analytical argument based on torque reduction by growing tails captures the essential physics and highlights the non‑equilibrium nature of the process. These results extend the classic Rouse description and open avenues for experimental validation and for incorporating additional interactions relevant to biological polymers.
Comments & Academic Discussion
Loading comments...
Leave a Comment