Single-molecule imaging of protein adsorption mechanisms to surfaces
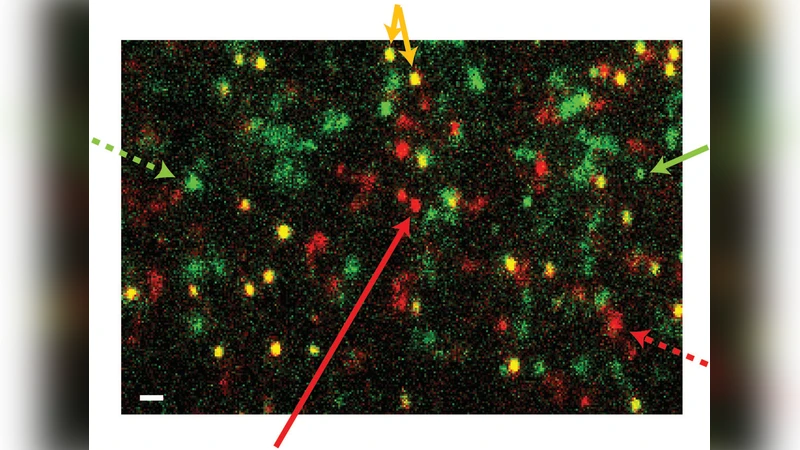
Protein-surface interactions cause the desirable effect of controlled protein adsorption onto biodevices as well as the undesirable effect of protein fouling. The key to controlling protein-surface adsorptions is to identify and quantify the main adsorption mechanisms: adsorptions that occur (1) while depositing a protein solution onto dry surfaces and (2) after the deposition onto wet surfaces. Bulk measurements cannot reveal the dynamic protein adsorption pathways and thus cannot differentiate between the two adsorption mechanisms. We imaged the interactions of single streptavidin molecules with hydrophobic fused-silica surfaces in real-time. We observed both adsorbed proteins on surfaces and diffusing proteins near surfaces and analyzed their adsorption kinetics. Our analysis shows that the protein solution deposition process is the primary mechanism of streptavidin adsorption onto surfaces at the sub-nanomolar to nanomolar protein concentrations. Furthermore, we found that hydrophilic fused-silica surfaces can prevent the adsorption of streptavidin molecules.
💡 Research Summary
This paper addresses the fundamental problem of protein adsorption on solid surfaces, a phenomenon that can be either beneficial (e.g., controlled immobilization on bio‑devices) or detrimental (e.g., fouling of medical implants). The authors argue that bulk techniques such as surface plasmon resonance or quartz‑crystal microbalance cannot resolve the two distinct pathways by which proteins reach a surface: (1) adsorption that occurs during the initial deposition of a protein solution onto a dry substrate, and (2) adsorption that takes place after the substrate is already wet. To dissect these pathways, they employ single‑molecule total internal reflection fluorescence (TIRF) microscopy to monitor individual streptavidin molecules labeled with Alexa 647 as they interact with fused‑silica substrates. Two surface chemistries are examined: a hydrophobic “fused‑silica” surface and a hydrophilic variant obtained by silanization that introduces surface hydroxyl groups.
Experimental design: streptavidin solutions at 0.01 nM, 0.1 nM, and 1 nM are introduced either (a) onto a freshly dried surface (the “deposition” condition) or (b) onto a surface that has already been wetted with buffer (the “post‑wet” condition). TIRF imaging captures both proteins that are freely diffusing within ~100 nm of the interface and those that become immobilized. Single‑particle tracking yields diffusion coefficients, residence times, and the temporal evolution of adsorption events.
Key findings: On the hydrophobic fused‑silica, the deposition condition generates a rapid burst of adsorption; roughly 80 % of all observed binding events at sub‑nanomolar to nanomolar concentrations arise during this initial contact. The average residence time of a bound streptavidin molecule under these conditions is ~30 seconds. In contrast, the post‑wet condition produces far fewer binding events (an order of magnitude lower) and most molecules simply diffuse near the surface without stable attachment. When the surface is rendered hydrophilic, both conditions show negligible adsorption; the few events detected are statistically indistinguishable from background. Concentration dependence reveals that at the lowest concentrations (≤0.1 nM) the deposition pathway dominates, whereas at higher concentrations (≥10 nM) the contribution of post‑wet adsorption becomes more noticeable.
Interpretation: The authors attribute the dominance of the deposition pathway to a transient concentration gradient that forms when a droplet contacts a dry surface. Rapid solvent evaporation and the lack of an existing hydration layer concentrate streptavidin at the interface, dramatically increasing the probability of direct collision and binding. A pre‑hydrated, hydrophilic surface, by contrast, maintains a stable water layer that screens electrostatic and hydrophobic interactions, thereby suppressing protein attachment.
Implications for device engineering: The study suggests that the most effective strategy to mitigate fouling is to avoid the “dry‑to‑wet” transition during manufacturing. Options include pre‑wetting the substrate, applying hydrophilic coatings (e.g., PEG, silanol‑rich layers), or controlling environmental humidity to limit rapid evaporation. The single‑molecule approach also provides a quantitative framework for comparing different protein–surface pairs, enabling predictive modeling of adsorption kinetics for a wide range of biomedical applications.
In summary, this work demonstrates that single‑molecule imaging can uniquely resolve the kinetic pathways of protein adsorption, identifies solution deposition as the primary mechanism of streptavidin binding on hydrophobic fused‑silica at low concentrations, and shows that hydrophilic surface modification effectively prevents such adsorption. These insights pave the way for more rational design of anti‑fouling surfaces and for the development of controlled protein immobilization strategies in bio‑engineering.
Comments & Academic Discussion
Loading comments...
Leave a Comment