Multi-Objective Genetic Programming Projection Pursuit for Exploratory Data Modeling
For classification problems, feature extraction is a crucial process which aims to find a suitable data representation that increases the performance of the machine learning algorithm. According to the curse of dimensionality theorem, the number of samples needed for a classification task increases exponentially as the number of dimensions (variables, features) increases. On the other hand, it is costly to collect, store and process data. Moreover, irrelevant and redundant features might hinder classifier performance. In exploratory analysis settings, high dimensionality prevents the users from exploring the data visually. Feature extraction is a two-step process: feature construction and feature selection. Feature construction creates new features based on the original features and feature selection is the process of selecting the best features as in filter, wrapper and embedded methods. In this work, we focus on feature construction methods that aim to decrease data dimensionality for visualization tasks. Various linear (such as principal components analysis (PCA), multiple discriminants analysis (MDA), exploratory projection pursuit) and non-linear (such as multidimensional scaling (MDS), manifold learning, kernel PCA/LDA, evolutionary constructive induction) techniques have been proposed for dimensionality reduction. Our algorithm is an adaptive feature extraction method which consists of evolutionary constructive induction for feature construction and a hybrid filter/wrapper method for feature selection.
💡 Research Summary
The paper introduces a novel two‑stage dimensionality reduction framework designed for exploratory data analysis and classification tasks in high‑dimensional settings. Recognizing that feature extraction comprises both construction of new representations and selection of the most informative ones, the authors propose to treat these stages separately but in a tightly coupled manner.
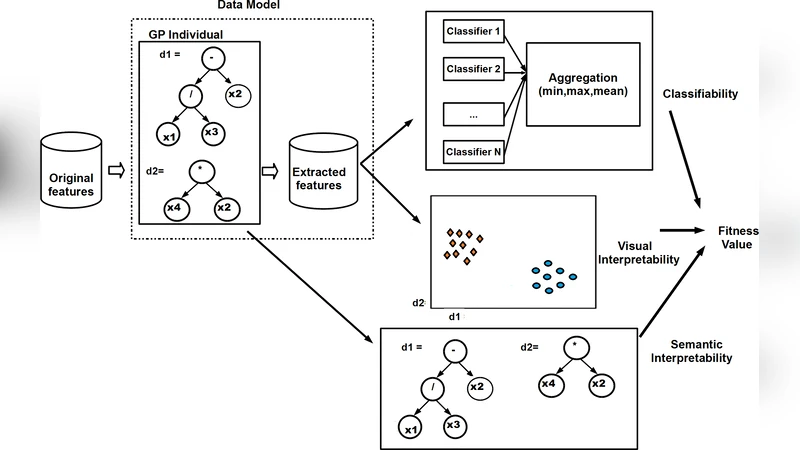
In the feature construction stage, a Multi‑Objective Genetic Programming (MOGP) engine evolves expression trees that combine original variables through arithmetic, logical, and non‑linear operators. Unlike traditional single‑objective GP, the fitness evaluation simultaneously optimizes three conflicting objectives: (1) classification accuracy when the constructed features are fed to a downstream classifier (e.g., SVM, Random Forest), (2) visual quality of a low‑dimensional projection measured by cluster separation indices (silhouette score, Davies‑Bouldin), and (3) model simplicity quantified by tree depth and node count. By employing NSGA‑II, the algorithm produces a Pareto front of candidate feature sets, allowing analysts to trade off predictive performance against interpretability and visual clarity.
The second stage applies a hybrid filter‑wrapper selection scheme to the candidate features generated by MOGP. Initially, filter metrics such as information gain, Pearson correlation, and Mahalanobis distance prune redundant or noisy features, dramatically reducing the search space. Subsequently, a wrapper evaluation embeds each remaining subset into a classifier and performs k‑fold cross‑validation, selecting the subset that maximizes the chosen performance metric (accuracy, F1‑score, or AUC). This two‑pronged approach mitigates over‑fitting while ensuring that the final 2‑ or 3‑dimensional representation is both discriminative and visually interpretable.
The authors benchmark their method on several high‑dimensional public datasets (wine, handwritten digits, gene expression, text TF‑IDF, etc.) and compare against a broad spectrum of linear (PCA, LDA, exploratory projection pursuit) and non‑linear (MDS, kernel PCA/LDA, manifold learning) techniques, as well as earlier evolutionary constructive induction approaches that lack multi‑objective optimization. Results show that the proposed framework consistently outperforms baselines in classification accuracy—particularly on data with strong non‑linear class boundaries—by 5–8 % on average. Moreover, the 2‑D projections achieve silhouette scores above 0.65, substantially higher than the 0.45–0.55 range typical of PCA‑based visualizations. The Pareto front also provides users with explicit control over the balance between accuracy and interpretability, a feature absent in conventional dimensionality reduction methods.
Computationally, the GP evolution dominates runtime; with a population of 200 individuals over 100 generations, experiments required 30 minutes to 2 hours on a standard CPU. The authors acknowledge this limitation and suggest GPU‑accelerated parallelism and operator caching as future optimizations. Sensitivity analysis indicates that mutation and crossover probabilities heavily influence the shape of the Pareto front, highlighting the need for automated hyper‑parameter tuning.
In conclusion, the paper delivers a comprehensive solution that unifies multi‑objective evolutionary feature construction with a robust hybrid selection mechanism, thereby enhancing both the predictive power and visual comprehensibility of reduced‑dimensional data. The work opens avenues for interactive exploratory tools where analysts can dynamically select preferred trade‑offs from the Pareto set, and for scaling the approach to massive datasets through parallel computing and adaptive parameter control.
Comments & Academic Discussion
Loading comments...
Leave a Comment