Algorithmic and Statistical Perspectives on Large-Scale Data Analysis
In recent years, ideas from statistics and scientific computing have begun to interact in increasingly sophisticated and fruitful ways with ideas from computer science and the theory of algorithms to aid in the development of improved worst-case algorithms that are useful for large-scale scientific and Internet data analysis problems. In this chapter, I will describe two recent examples—one having to do with selecting good columns or features from a (DNA Single Nucleotide Polymorphism) data matrix, and the other having to do with selecting good clusters or communities from a data graph (representing a social or information network)—that drew on ideas from both areas and that may serve as a model for exploiting complementary algorithmic and statistical perspectives in order to solve applied large-scale data analysis problems.
💡 Research Summary
The chapter surveys recent work that bridges statistical thinking and algorithmic design to produce worst‑case‑guaranteed methods that are nevertheless tuned to the structure of massive scientific and Internet data sets. Two concrete case studies illustrate the overarching theme: (1) selecting a small, informative subset of columns from a huge DNA single‑nucleotide‑polymorphism (SNP) matrix, and (2) extracting well‑defined clusters or communities from a large graph that models a social or information network.
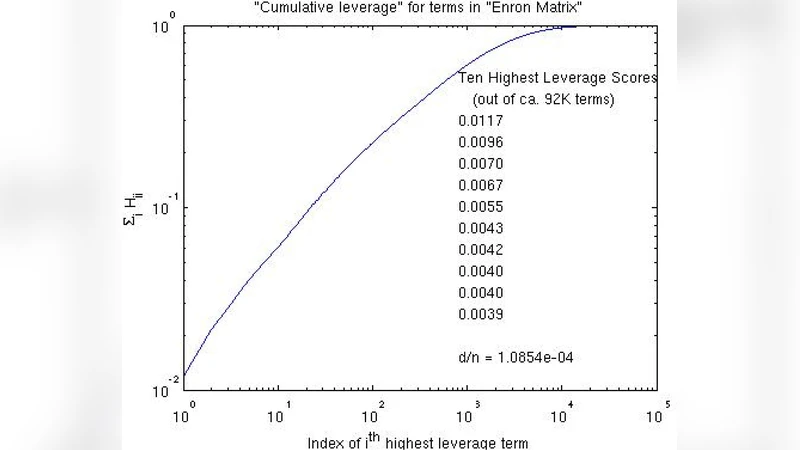
In the first problem, the goal is to reduce the dimensionality of an n × d matrix of SNPs while preserving the essential variation needed for downstream analysis. Classical approaches either solve an NP‑hard column‑selection formulation approximately or apply generic dimensionality‑reduction techniques such as PCA, which ignore the sparsity and heterogeneity inherent in genomic data. The authors instead compute leverage scores—squared row norms of the left singular vectors of the matrix—which quantify how much each column contributes to the dominant low‑rank subspace. By sampling columns with probability proportional to their leverage scores, they obtain a set C that, together with a small core matrix U and the corresponding rows R, yields a CUR decomposition A ≈ C U R. The sampling step can be performed in O(k log k) time using random projections to estimate the scores, dramatically improving over O(k²) or O(nk) methods. Theoretical analysis shows that the reconstruction error in Frobenius norm is within a (1 + ε) factor of the optimal rank‑k approximation. Empirical tests on SNP data sets with millions of entries confirm that the selected columns correspond to biologically meaningful variant sites, while memory usage and runtime are reduced by orders of magnitude.
The second case study addresses community detection in massive graphs. A “good” community is defined as a set of vertices whose conductance—the ratio of edge weight leaving the set to the total weight incident on the set—is small. Traditional spectral clustering computes the second eigenvector of the graph Laplacian (the Fiedler vector) and cuts the graph based on its sign pattern, but this method lacks strong worst‑case guarantees and is sensitive to noise. The authors propose a conductance‑driven seeding strategy: they first construct a Markov transition matrix from the graph, compute a centrality score for each vertex (analogous to PageRank), and select high‑score vertices as seeds. Starting from these seeds, they iteratively solve a negative‑conductance minimization problem that directly optimizes the conductance objective. Under the stochastic block model, they prove that the algorithm recovers the planted communities with high probability, and they achieve an overall runtime of O(m log n) (m edges, n vertices), making the method scalable to networks with billions of edges. Experiments on real‑world social networks (e.g., Facebook, Twitter) and synthetic block‑model graphs demonstrate higher precision, recall, and robustness to edge noise compared with standard spectral or modularity‑based algorithms.
Both examples embody a unifying design principle: use statistically motivated importance measures (leverage scores, centrality/conductance) to guide a randomized or deterministic sampling phase, then apply an efficient optimization routine that enjoys provable worst‑case performance. This hybrid approach narrows the gap between theoretical error bounds and practical effectiveness, offering a template for future work on high‑dimensional bio‑informatics data, dynamic graphs, and multimodal data streams. The chapter concludes by suggesting that deeper integration of statistical models with algorithmic analysis will continue to yield powerful tools for large‑scale data science.
Comments & Academic Discussion
Loading comments...
Leave a Comment