Induced fit, conformational selection and independent dynamic segments: an extended view of binding events
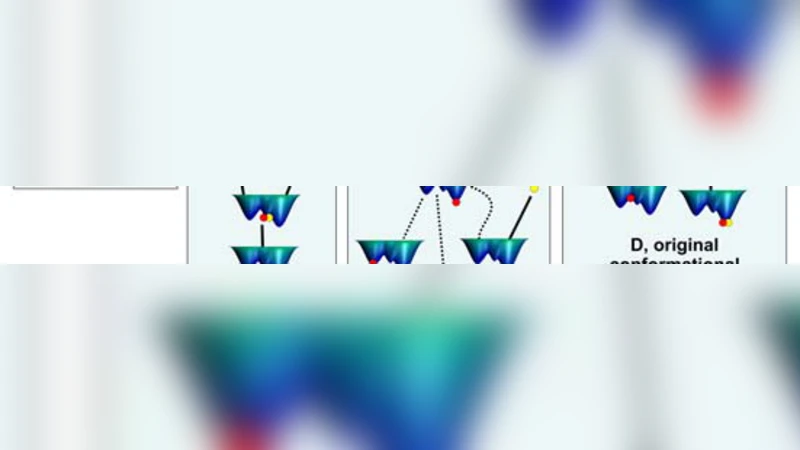
Single molecule and NMR measurements of protein dynamics increasingly uncover the complexity of binding scenarios. Here we describe an extended conformational selection model which embraces a repertoire of selection and adjustment processes. Induced fit can be viewed as a subset of this repertoire, whose contribution is affected by the bond-types stabilizing the interaction and the differences between the interacting partners. We argue that protein segments whose dynamics are distinct from the rest of the protein (‘discrete breathers’) can govern conformational transitions and allosteric propagation that accompany binding processes, and as such may be more sensitive to mutational events. Additionally, we highlight the dynamic complexity of binding scenarios as they relate to events such as aggregation and signalling, and the crowded cellular environment.
💡 Research Summary
The authors present an expanded view of protein‑binding mechanisms that integrates the classic conformational‑selection and induced‑fit paradigms into a unified, multi‑step framework. Starting from single‑molecule and NMR observations, they argue that binding is rarely a single, static event; instead it proceeds through a sequence of “selection → adjustment → transition” stages. In the selection phase, an ensemble of pre‑existing conformers is sampled, and the ligand preferentially binds the most compatible substate. Immediately after binding, a rapid local rearrangement—termed “adjustment”—occurs, which corresponds to the traditional induced‑fit component but is framed here as a transient, cooperative response rather than a separate mechanism.
A central novelty of the paper is the introduction of “independent dynamic segments,” also called discrete breathers, which are protein regions whose intrinsic motions are decoupled from the rest of the structure. These segments act as localized energy concentrators, lowering activation barriers for conformational transitions and providing a directed pathway for the structural shift from the selected to the final bound state. Because they mediate energy flow, discrete breathers are especially sensitive to point mutations; alterations in these regions can disproportionately affect binding kinetics, allosteric signaling, and overall protein stability. The authors support this claim with mutational data showing a higher incidence of disease‑associated variants near identified dynamic segments.
The paper also situates the model within the crowded cellular milieu. Macromolecular crowding alters solvent properties and imposes volume‑exclusion forces, reshaping the free‑energy landscape of proteins. Under such conditions, both conformational selection and induced‑fit contributions are amplified, and the role of discrete breathers becomes more pronounced. For example, crowding‑induced compression can trigger the activation of a dynamic segment, thereby accelerating the transition step and enabling rapid signaling responses.
Beyond basic binding, the authors discuss how the same dynamic principles influence protein aggregation and signal transduction. In early aggregation, exposed dynamic segments can promote nonspecific contacts that seed oligomer formation. Conversely, in signaling pathways, discrete breathers can act as molecular switches, selectively enhancing interactions with specific partners while suppressing off‑target binding. This dual functionality illustrates how a single protein can adopt distinct functional outcomes by modulating the same set of dynamic elements.
In summary, the manuscript proposes a comprehensive, energetically grounded model of protein binding that moves beyond a binary classification of conformational selection versus induced fit. By emphasizing the pivotal role of independent dynamic segments, the authors provide a mechanistic explanation for mutation sensitivity, allosteric propagation, and environmental modulation. This extended framework has practical implications for drug design—where targeting dynamic hotspots may yield higher specificity—and for interpreting the impact of genetic variation on protein function in vivo.
Comments & Academic Discussion
Loading comments...
Leave a Comment