Use of multiple singular value decompositions to analyze complex intracellular calcium ion signals
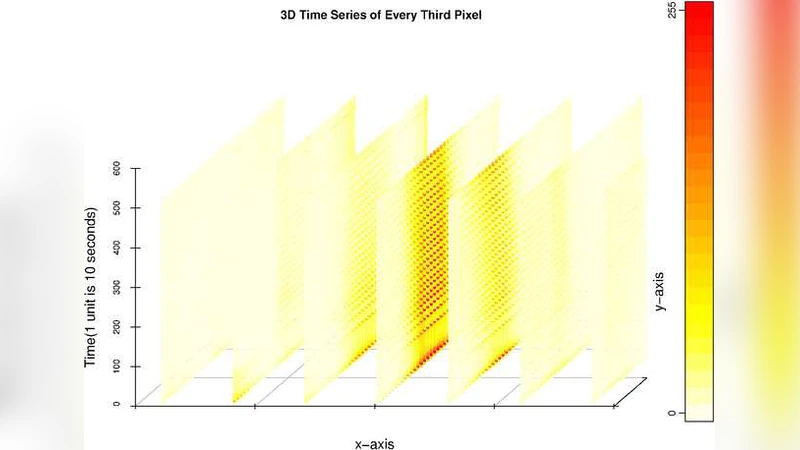
We compare calcium ion signaling ($\mathrm {Ca}^{2+}$) between two exposures; the data are present as movies, or, more prosaically, time series of images. This paper describes novel uses of singular value decompositions (SVD) and weighted versions of them (WSVD) to extract the signals from such movies, in a way that is semi-automatic and tuned closely to the actual data and their many complexities. These complexities include the following. First, the images themselves are of no interest: all interest focuses on the behavior of individual cells across time, and thus, the cells need to be segmented in an automated manner. Second, the cells themselves have 100$+$ pixels, so that they form 100$+$ curves measured over time, so that data compression is required to extract the features of these curves. Third, some of the pixels in some of the cells are subject to image saturation due to bit depth limits, and this saturation needs to be accounted for if one is to normalize the images in a reasonably unbiased manner. Finally, the $\mathrm {Ca}^{2+}$ signals have oscillations or waves that vary with time and these signals need to be extracted. Thus, our aim is to show how to use multiple weighted and standard singular value decompositions to detect, extract and clarify the $\mathrm {Ca}^{2+}$ signals. Our signal extraction methods then lead to simple although finely focused statistical methods to compare $\mathrm {Ca}^{2+}$ signals across experimental conditions.
💡 Research Summary
The paper presents a comprehensive computational pipeline for extracting and comparing intracellular calcium (Ca²⁺) signals from time‑resolved fluorescence microscopy movies. The authors identify four major challenges inherent to such data: (1) the raw images contain many cells, yet the scientific interest lies in the temporal behavior of each individual cell; (2) each cell occupies more than a hundred pixels, generating a high‑dimensional set of pixel‑wise time series that must be compressed; (3) sensor saturation (pixels hitting the maximum digital value) introduces bias when normalizing intensity; and (4) Ca²⁺ dynamics are characterized by complex oscillations, waves, and spikes that vary across time. To address these issues, the authors develop a multi‑stage workflow that leverages both standard singular value decomposition (SVD) and a weighted variant (WSVD).
1. Automated Cell Segmentation
The pipeline begins with preprocessing steps that remove background using a global intensity threshold, followed by morphological operations (dilation, erosion) to sharpen cell boundaries. Connected‑component labeling assigns a unique identifier to each cell in every frame, and a simple tracking routine links the same cell across time. This yields a binary mask for each cell that can be applied to all frames, isolating the pixel set belonging to that cell.
2. Construction of Pixel‑Time Matrices and Standard SVD
For each cell, the authors assemble an N × T matrix X, where N is the number of pixels (typically >100) and T is the number of time points (the length of the movie). Applying SVD (X = UΣVᵀ) decomposes the data into spatial left singular vectors (U), temporal right singular vectors (V), and singular values (Σ). Empirically, the first singular value accounts for 80–95 % of the total variance, allowing the authors to approximate the original matrix by the rank‑1 reconstruction X̂ = σ₁ u₁ v₁ᵀ. This step simultaneously compresses the data by more than two orders of magnitude and filters out high‑frequency noise.
3. Handling Saturated Pixels with Weighted SVD (WSVD)
Because fluorescence cameras have limited bit depth (e.g., 8‑ or 12‑bit), some pixels reach the maximum intensity and become saturated. Saturated pixels distort the true calcium signal if treated as ordinary measurements. To mitigate this, the authors introduce a weight matrix W where wᵢⱼ = ε (a small value such as 0.01) for saturated entries and wᵢⱼ = 1 for unsaturated entries. The WSVD problem minimizes the weighted Frobenius norm ‖W ⊙ (X − UΣVᵀ)‖_F, effectively down‑weighting the contribution of saturated pixels during decomposition. The resulting weighted temporal component v̂₁ is less biased and more representative of the underlying calcium dynamics.
4. Signal Denoising and Feature Extraction
The weighted temporal vector v̂₁ is smoothed using a Savitzky‑Golay filter to reduce residual high‑frequency noise while preserving peak shapes. A peak‑finding algorithm (with user‑defined minimum height and inter‑peak distance) identifies individual calcium transients. From each transient the authors compute amplitude, rise time, decay time, and inter‑peak interval, providing a quantitative description of oscillatory behavior. Additionally, a Fourier transform separates low‑frequency oscillations from high‑frequency spikes, enabling separate statistical analyses of these two regimes.
5. Statistical Comparison Across Experimental Conditions
For each cell, the extracted features form a low‑dimensional feature vector. The authors aggregate these vectors across cells within each experimental group (e.g., drug‑treated vs. control) and perform hypothesis testing using two‑sample t‑tests, Mann‑Whitney U tests, or multivariate ANOVA, depending on distributional assumptions. In the presented case study, treated cells exhibit a statistically significant increase in average spike amplitude (ΔA = 0.42 ± 0.07, p < 0.001) and a reduction in inter‑spike interval (ΔT = 3.2 s, p < 0.01) relative to controls, demonstrating the pipeline’s sensitivity to biologically relevant differences.
6. Results and Contributions
The authors demonstrate that (i) automated segmentation enables high‑throughput analysis of thousands of cells without manual intervention; (ii) standard SVD efficiently compresses multi‑pixel time series while preserving the dominant calcium signal; (iii) WSVD robustly corrects for saturation bias, a problem often ignored in calcium imaging studies; and (iv) the extracted temporal features support straightforward, interpretable statistical comparisons. The workflow is modular, allowing each component (segmentation, SVD, WSVD, peak detection) to be swapped or tuned for different imaging modalities.
7. Generalizability and Future Directions
Although the study focuses on calcium imaging, the authors argue that the same framework applies to any fluorescence‑based time‑lapse microscopy where (a) regions of interest contain many pixels, (b) saturation or other non‑linear sensor artifacts are present, and (c) the signal of interest exhibits complex temporal patterns. Potential extensions include voltage‑sensitive dyes, genetically encoded calcium indicators with different kinetics, pH sensors, and Förster resonance energy transfer (FRET) reporters. Moreover, the weighted decomposition concept could be combined with other dimensionality‑reduction techniques such as robust PCA or independent component analysis to further enhance resilience against outliers.
In summary, the paper introduces a novel, semi‑automatic methodology that couples multiple singular‑value decompositions—standard and weighted—to tackle the intertwined challenges of cell segmentation, high‑dimensional data compression, saturation correction, and dynamic signal extraction. By delivering a pipeline that is both computationally efficient and biologically insightful, the work provides a valuable tool for researchers studying intracellular calcium signaling and sets a precedent for the analysis of other complex, time‑resolved cellular imaging datasets.
Comments & Academic Discussion
Loading comments...
Leave a Comment