How is the Monoclonal Antibodies Kinetic Affected by Changes of Their Physical Parameters?
The study of monoclonal antibodies (MAb) is a field of great interest to science medicine, for example, anti-TNF agents (infliximab and adalimumab) represent an important tool for the management of autoimmune and inflammatory disorders. In this work we focus on the physical description of the transport kinetics of MAb in a fluid with laminar flow and parabolic profile. To simulate the kinetics of the MAb, standard equations were solved numerically (using The Verlet algorithm) to calculate the motion of a particle with a spherically symmetric inside of parabolic laminar flow, in order to find the time evolution of the antibody velocity in blood plasma in function of the increase of the radius, mass and density of the MAb, and the fluid pressure in blood vessels. In the case of we fixed the value of the antibody density, their kinetics increased when the pressure in the vessels increased. When we fixed the pressure in the vessels we found: if we reduce the antibody radius their kinetics increased, and when we increase antibody density we found that their kinetics also increased.
💡 Research Summary
The manuscript investigates how the physical parameters of monoclonal antibodies (MAbs) – specifically radius, mass (or equivalently density), and the surrounding fluid pressure – influence their transport kinetics in blood plasma under laminar, parabolic flow conditions. The authors model a blood vessel as a straight cylindrical pipe with a fully developed parabolic velocity profile, (u(r)=U_{\max}(1-(r/R)^2)), where (R) is the vessel radius. An antibody is treated as a rigid, spherical particle characterized by radius (a), density (\rho_p), and mass (m = \frac{4}{3}\pi a^3\rho_p). The forces acting on the particle are (i) the pressure‑driven body force (-\nabla P,V) (with (V) the particle volume), (ii) Stokes drag (F_d = 6\pi\mu a (v_f - v_p)) where (\mu) is plasma viscosity, and (iii) inertial terms from the particle’s own mass.
To solve the particle’s equation of motion, the authors employ the Verlet integration scheme, a time‑stepping algorithm known for its good energy‑conserving properties in conservative systems. Initial conditions place the particle at rest at the inlet of the pipe; the simulation then advances the particle’s position and velocity under the combined forces. Two families of parametric sweeps are performed: (1) density (\rho_p) is held constant while the pressure gradient (\nabla P) is varied, and (2) the pressure gradient is fixed while radius (a) and density (\rho_p) are independently altered.
Key findings are as follows:
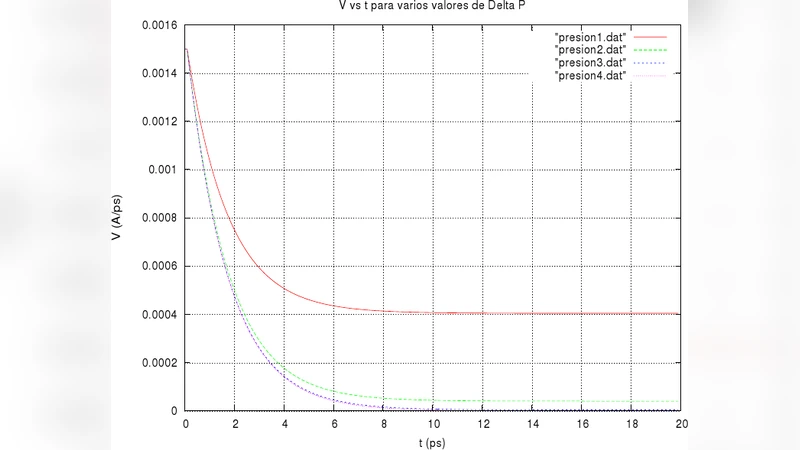
- Pressure dependence: With a fixed particle density, increasing the pressure gradient leads to a monotonic increase in both the instantaneous velocity and the overall acceleration of the antibody. This result aligns with the intuitive expectation that a larger pressure drop supplies a greater driving force.
- Size dependence: At a constant pressure gradient, decreasing the particle radius markedly speeds up the particle. Because Stokes drag scales linearly with radius, smaller particles experience less viscous resistance and thus follow the fluid streamlines more closely.
- Density dependence: Holding the pressure gradient constant, raising the particle density also enhances its kinetic response. A higher density raises the particle’s inertia, allowing it to maintain momentum against viscous drag and to respond more rapidly to the pressure‑driven force.
The authors interpret these outcomes as guidance for antibody engineering: a design that minimizes hydrodynamic radius while maximizing effective density (for example, by conjugating dense nanomaterials) could improve delivery speed in high‑pressure vascular regions.
Despite its conceptual appeal, the study has several methodological limitations. First, blood is treated as a Newtonian fluid, whereas real plasma exhibits shear‑thinning behavior and the presence of red blood cells introduces non‑laminar micro‑scale disturbances. Second, the use of Verlet integration, while advantageous for conservative dynamics, may be suboptimal for highly dissipative Stokes‑dominated regimes; alternative schemes (e.g., implicit Euler or Runge‑Kutta with adaptive stepping) could provide better stability. Third, the manuscript lacks a systematic convergence analysis (time‑step sensitivity, spatial discretization) and does not report quantitative validation against experimental data such as microfluidic measurements of antibody transport. Fourth, Brownian motion, which becomes significant for particles below ~100 nm, is omitted, potentially over‑estimating the deterministic velocity for very small antibodies or antibody‑nanoparticle conjugates. Finally, the physiological range of pressures (≈ 80–120 mmHg) and flow rates (≈ 0.1–0.5 m s⁻¹) is not explicitly linked to the simulation parameters, making it difficult to assess the real‑world relevance of the reported trends.
Future work should address these gaps by incorporating non‑Newtonian rheology, vessel wall compliance, and pulsatile pressure waveforms into a full computational fluid dynamics (CFD) framework. Coupling the particle dynamics with stochastic diffusion terms would capture the transition from deterministic advection to diffusion‑dominated regimes. Experimental validation using microfluidic channels that mimic arterial shear rates, together with high‑speed imaging of fluorescently labeled antibodies of varying size and density, would provide essential benchmarks. Ultimately, extending the model to include antigen‑binding kinetics and immune complex formation would enable a comprehensive pharmacokinetic‑pharmacodynamic (PK‑PD) description, turning the present theoretical insights into actionable design rules for next‑generation therapeutic antibodies.
Comments & Academic Discussion
Loading comments...
Leave a Comment