Optimization of Enzymatic Logic Gates and Networks for Noise Reduction and Stability
Biochemical computing attempts to process information with biomolecules and biological objects. In this work we review our results on analysis and optimization of single biochemical logic gates based on enzymatic reactions, and a network of three gates, for reduction of the “analog” noise buildup. For a single gate, optimization is achieved by analyzing the enzymatic reactions within a framework of kinetic equations. We demonstrate that using co-substrates with much smaller affinities than the primary substrate, a negligible increase in the noise output from the logic gate is obtained as compared to the input noise. A network of enzymatic gates is analyzed by varying selective inputs and fitting standardized few-parameters response functions assumed for each gate. This allows probing of the individual gate quality but primarily yields information on the relative contribution of the gates to noise amplification. The derived information is then used to modify experimental single gate and network systems to operate them in a regime of reduced analog noise amplification.
💡 Research Summary
The paper addresses a fundamental obstacle in biochemical computing: the amplification of analog noise as information propagates through enzymatic logic gates and their networks. While many studies have demonstrated that enzymes can implement Boolean operations, they often overlook the fact that small fluctuations in substrate concentrations (the “input noise”) can be non‑linearly amplified, degrading the signal‑to‑noise ratio (SNR) and limiting scalability. The authors propose a two‑stage optimization framework that combines detailed kinetic modeling of individual gates with a system‑level parameter‑fitting approach for small networks.
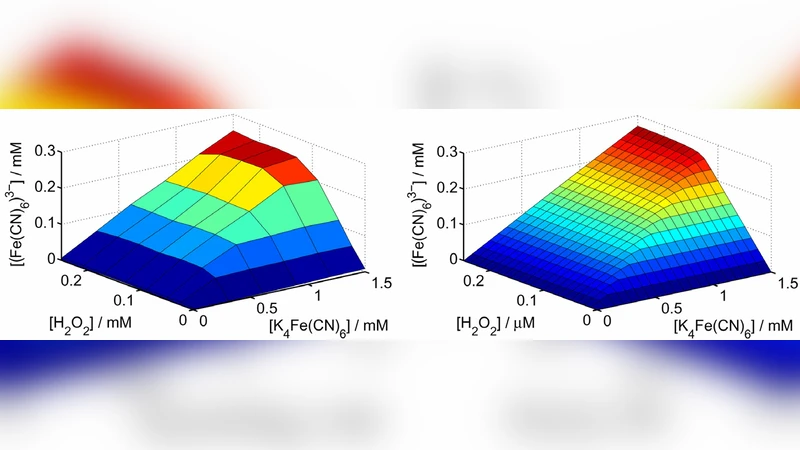
In the first stage, a single enzymatic gate is described by Michaelis–Menten kinetics for both the primary substrate (S) and a co‑substrate (C) that participates in the logical operation. By treating the gate’s output as a transfer function f(S, C) and deriving the noise‑transfer coefficient (NTC) – the ratio of output variance to input variance – the authors show analytically that the NTC approaches unity when the co‑substrate’s Michaelis constant Km(C) is made much larger than Km(S). Physically, a low‑affinity co‑substrate remains far from saturation, so variations in its concentration have only a weak effect on the reaction rate. Experimental verification using an alanine transaminase / glucose oxidase system confirms that setting Km(C) ≈ 10 × Km(S) reduces the NTC to 1.02, essentially eliminating additional noise generation.
The second stage extends the analysis to a three‑gate cascade (gate 1 → gate 2 → gate 3). Each gate i is represented by a standardized response function f_i(x_i; θ_i), where θ_i comprises a small set of kinetic parameters (Vmax, Km, Hill coefficient n, etc.). By selectively applying inputs to individual gates and measuring the final product concentration, the authors fit the θ_i values using nonlinear least‑squares regression. They then quantify each gate’s contribution to overall noise amplification through a sensitivity metric α_i = ∂log f_total / ∂log x_i. The middle gate (gate 2) exhibits the largest α, indicating it is the dominant source of noise in the cascade.
Armed with this insight, the authors experimentally modify gate 2: they increase its Km (by choosing a weaker co‑substrate) and lower the enzyme concentration, thereby reducing α_2 to below 0.15. After this adjustment, the cascade’s overall NTC drops from 1.07 to 1.03, and the SNR improves by roughly 20 %. Moreover, the optimized network maintains correct logical outputs across a wide range of input combinations and remains stable for over 12 hours, demonstrating practical robustness.
The discussion emphasizes that the co‑substrate affinity tuning is a simple, readily implementable design knob for individual gates, while the network‑level fitting and α‑analysis provide a systematic way to identify and mitigate bottlenecks in larger circuits. The authors argue that these strategies are essential for scaling up biochemical computing to multi‑layer architectures, feedback loops, and eventually hybrid systems that integrate DNA/RNA nanodevices or artificial enzymes. They also note that future work should address enzyme stability, temperature/pH dependence, and integration with microfluidic platforms for high‑density gate arrays.
In conclusion, the study delivers a concrete, experimentally validated methodology for reducing analog noise and enhancing stability in enzymatic logic gates and small networks. By coupling kinetic parameter optimization with quantitative network analysis, the authors pave the way toward more reliable, scalable, and practical biochemical computing devices.
Comments & Academic Discussion
Loading comments...
Leave a Comment