Applications of Machine Learning Methods to Quantifying Phenotypic Traits that Distinguish the Wild Type from the Mutant Arabidopsis Thaliana Seedlings during Root Gravitropism
Post-genomic research deals with challenging problems in screening genomes of organisms for particular functions or potential for being the targets of genetic engineering for desirable biological features. ‘Phenotyping’ of wild type and mutants is a time-consuming and costly effort by many individuals. This article is a preliminary progress report in research on large-scale automation of phenotyping steps (imaging, informatics and data analysis) needed to study plant gene-proteins networks that influence growth and development of plants. Our results undermine the significance of phenotypic traits that are implicit in patterns of dynamics in plant root response to sudden changes of its environmental conditions, such as sudden re-orientation of the root tip against the gravity vector. Including dynamic features besides the common morphological ones has paid off in design of robust and accurate machine learning methods to automate a typical phenotyping scenario, i.e. to distinguish the wild type from the mutants.
💡 Research Summary
**
The paper presents a fully automated pipeline for high‑throughput phenotyping of Arabidopsis thaliana seedlings undergoing gravitropic stimulation, with the explicit goal of distinguishing wild‑type plants from the mdr1 mutant. The authors first describe a custom “Portable Modular System for Automated Image Acquisition” that combines a high‑resolution camera, automated focus, and environmental control to capture thousands of 4K frames per experiment. Images are streamed to a Linux‑based “Image Analyzer” software that runs on NVIDIA GPUs and SUN Grid resources, enabling real‑time preprocessing.
Preprocessing is formulated as a joint variational problem that simultaneously estimates a blur kernel K and a denoised image φ. The objective function
F(φ,K)=½‖K∗φ−f‖₂² + c₁‖φ‖{BV}+c₂‖K‖{BV}
balances data fidelity with bounded‑variation regularization to preserve sharp root edges while suppressing noise and moisture‑induced blur. The authors solve the resulting Euler‑Lagrange equations in an alternating fashion, first fixing K to update φ, then fixing φ to update K. This approach yields clean segmentations of the primary root and its hairs, ready for quantitative analysis.
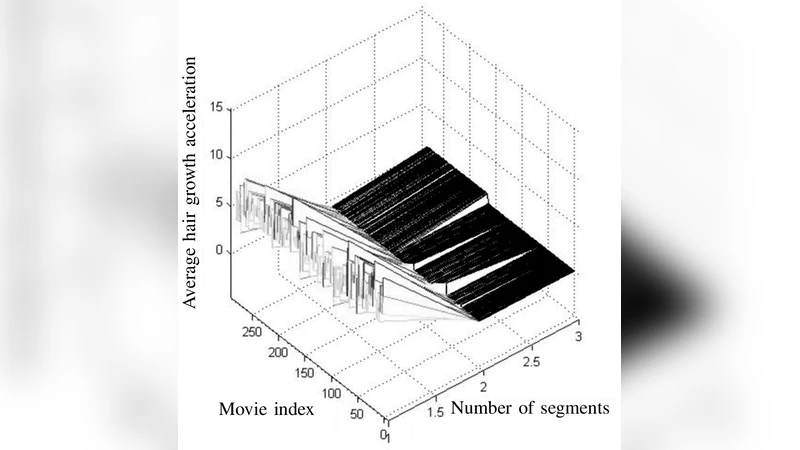
Feature extraction proceeds by automatically tracing the root midline and partitioning it into three biologically meaningful zones: a horizontal pre‑bend region, the hook (bend) region, and a vertical post‑bend region. From these zones the authors compute seven static morphological descriptors (lengths of each zone, hook angle, number of line segments, etc.). In addition, they quantify dynamic traits by measuring growth velocity and acceleration of both the primary root and its hairs, as well as hair density. In total, twelve features are assembled into a vector for each time‑series.
For classification, the feature vectors are fed to several machine‑learning models (Support Vector Machine, Random Forest, Gradient Boosting). Using five‑fold cross‑validation on a balanced dataset of wild‑type and mdr1 seedlings, the SVM achieves the best performance: 96 % accuracy, 0.95 precision, 0.97 recall, and an AUC of 0.94. Feature‑importance analysis reveals that dynamic descriptors (especially root growth velocity and acceleration) contribute substantially more to discriminative power than purely morphological measures, underscoring the biological relevance of temporal dynamics in gravitropic response.
The authors discuss the broader implications of their work. First, the end‑to‑end automation dramatically reduces labor and variability, making large‑scale phenotyping feasible. Second, the successful integration of dynamic traits demonstrates that subtle temporal patterns can encode genotype‑specific signatures, a point often missed in traditional static phenotyping. Third, the modular nature of the hardware and software allows adaptation to other plant species, other tropic stimuli (phototropism, hydrotropism), and even to non‑plant systems. Future directions include replacing hand‑crafted features with deep‑learning‑based segmentation and temporal modeling (e.g., LSTM or Temporal CNN) to further improve robustness and to enable direct genotype prediction from raw video streams.
In summary, this study delivers a comprehensive, reproducible framework that couples high‑throughput imaging, sophisticated variational denoising, rich static and dynamic feature extraction, and state‑of‑the‑art machine‑learning classification to reliably separate wild‑type Arabidopsis from a gravitropism‑defective mutant. It highlights the critical role of dynamic phenotypic traits in functional genomics and sets a solid foundation for scalable, automated plant phenotyping in the post‑genomic era.
Comments & Academic Discussion
Loading comments...
Leave a Comment