Network motifs come in sets: correlations in the randomization process
The identification of motifs–subgraphs that appear significantly more often in a particular network than in an ensemble of randomized networks–has become a ubiquitous method for uncovering potentially important subunits within networks drawn from a wide variety of fields. We find that the most common algorithms used to generate the ensemble from the real network change subgraph counts in a highly correlated manner, so that one subgraph’s status as a motif may not be independent from the statuses of the other subgraphs. We demonstrate this effect for the problem of 3- and 4-node motif identification in the transcriptional regulatory networks of E. coli and S. cerevisiae in which randomized networks are generated via an edge-swapping algorithm (Milo et al., Science 298:824, 2002). We show that correlations among 3-node subgraphs are easily interpreted, and we present an information-theoretic tool that may be used to identify correlations among subgraphs of any size.
💡 Research Summary
The paper revisits a foundational assumption in network‑motif analysis: that each small subgraph can be evaluated independently for statistical over‑representation. Motif detection typically proceeds by comparing the frequency of each subgraph in a real network with its frequency in an ensemble of randomized networks that preserve certain global properties, most commonly the degree sequence. The standard way to generate such an ensemble is the edge‑swap (or degree‑preserving rewiring) algorithm introduced by Milo et al. (2002). In this algorithm two directed edges are selected at random and their endpoints are exchanged, thereby preserving the in‑ and out‑degree of every node while scrambling the wiring pattern.
The authors focus on transcriptional regulatory networks of Escherichia coli and Saccharomyces cerevisiae. They enumerate all possible 3‑node (13 directed patterns) and 4‑node (199 directed patterns) subgraphs, count their occurrences in the original networks, and generate 10,000+ randomized copies using the edge‑swap method. For each subgraph they compute a Z‑score and a p‑value to assess “motifness”. The key observation is that the edge‑swap process does not perturb subgraph counts independently. Instead, many pairs of subgraphs change in a tightly coupled fashion: a decrease in one is almost always accompanied by a corresponding increase in another. For example, in the 3‑node case the feed‑forward loop (FFL) and the feed‑back loop (FBL) exhibit an almost one‑to‑one trade‑off during each swap. Similar coupled transformations appear among 4‑node patterns, such as between a fully connected 4‑node clique and a configuration consisting of two overlapping 3‑node FFLs.
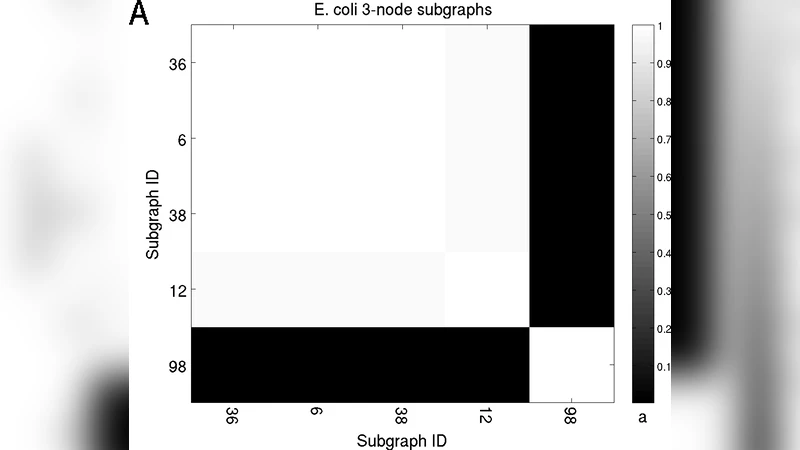
To quantify these dependencies the authors introduce an information‑theoretic measure: the mutual information (I(A;B)=\sum_{a,b} P(a,b)\log\frac{P(a,b)}{P(a)P(b)}) between the count distributions of two subgraphs (A) and (B) across the randomized ensemble. High mutual information indicates that the two counts co‑vary more than expected under independence. Empirically, several 3‑node and 4‑node pairs show substantial mutual information, confirming that the randomization process creates systematic correlations.
The paper also explores alternative randomization schemes. When the same networks are randomized using a multi‑step degree‑preserving rewiring that selects larger edge‑sets or applies Metropolis‑Hastings acceptance criteria, the observed correlations weaken markedly. This suggests that the strength of the artifact is algorithm‑specific rather than an unavoidable consequence of degree preservation.
The implications are profound for motif research. Because the statistical significance of a subgraph is traditionally assessed in isolation, any hidden correlation can inflate or deflate its Z‑score depending on the behavior of its partners. Consequently, a subgraph flagged as a motif may owe its status partly to the way the random ensemble was constructed, not solely to genuine biological design. The authors propose two practical remedies: (1) compute mutual information for all subgraph pairs and use it as a correction factor or as a filter to identify “dependent” motifs; (2) adopt multivariate statistical models (e.g., multiple regression or Bayesian hierarchical models) that treat the full vector of subgraph counts jointly, thereby accounting for covariance structure.
Biologically, the study does not dispute that certain motifs such as the FFL play functional roles in transcriptional regulation. Rather, it cautions that the statistical evidence for over‑representation must be interpreted in light of the randomization method. Researchers should report the randomization algorithm in detail, test multiple algorithms when feasible, and consider the possibility that observed motif enrichment could be an artifact of correlated count changes.
In conclusion, the paper demonstrates that the widely used edge‑swap randomization introduces strong, systematic correlations among subgraph frequencies, undermining the independence assumption underlying traditional motif detection. By employing mutual information as a diagnostic tool and advocating for more sophisticated randomization or statistical correction strategies, the authors provide a clear pathway to more reliable identification of truly significant network motifs.
Comments & Academic Discussion
Loading comments...
Leave a Comment