Uniformly curated signaling pathways reveal tissue-specific cross-talks and support drug target discovery
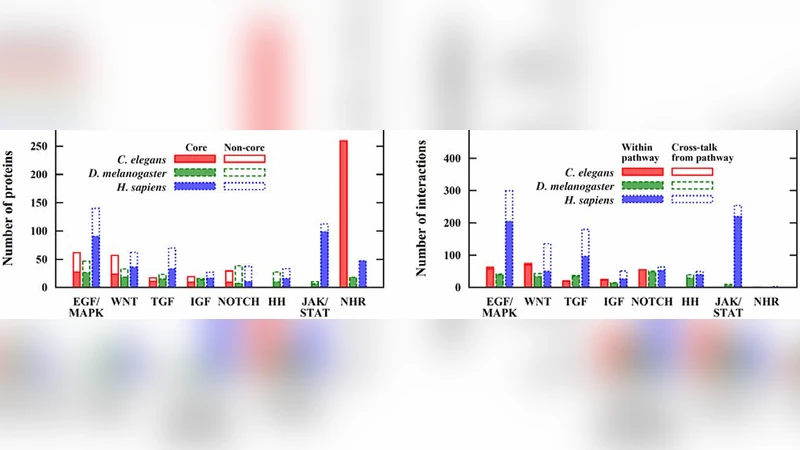
Motivation: Signaling pathways control a large variety of cellular processes. However, currently, even within the same database signaling pathways are often curated at different levels of detail. This makes comparative and cross-talk analyses difficult. Results: We present SignaLink, a database containing 8 major signaling pathways from Caenorhabditis elegans, Drosophila melanogaster, and humans. Based on 170 review and approx. 800 research articles, we have compiled pathways with semi-automatic searches and uniform, well-documented curation rules. We found that in humans any two of the 8 pathways can cross-talk. We quantified the possible tissue- and cancer-specific activity of cross-talks and found pathway-specific expression profiles. In addition, we identified 327 proteins relevant for drug target discovery. Conclusions: We provide a novel resource for comparative and cross-talk analyses of signaling pathways. The identified multi-pathway and tissue-specific cross-talks contribute to the understanding of the signaling complexity in health and disease and underscore its importance in network-based drug target selection. Availability: http://SignaLink.org
💡 Research Summary
Signaling pathways orchestrate a vast array of cellular processes, yet existing pathway databases often curate the same pathway at different levels of detail, hampering comparative and cross‑talk analyses. To address this, the authors built SignaLink, a uniformly curated resource that integrates eight major signaling pathways—such as EGFR, TGF‑β, Wnt, Notch, Hedgehog, JAK/STAT, MAPK, and PI3K/AKT—from three model organisms: Caenorhabditis elegans, Drosophila melanogaster, and Homo sapiens. The curation pipeline combined semi‑automatic literature mining with strict, documented rules that define five core interaction types (protein‑protein binding, phosphorylation/dephosphorylation, transcription factor activation, ligand‑receptor binding, and complex formation). Over 170 review articles and roughly 800 primary research papers were examined, resulting in a consistent representation of nodes (proteins or complexes) and edges (interactions) together with provenance information.
In the human network, the authors systematically examined all 28 possible pathway pairs and discovered that 27 pairs (96 %) share at least one cross‑talk protein, indicating that any two of the eight pathways can potentially interact. To quantify these interactions, they introduced a “cross‑talk strength” metric that accounts for the centrality of the shared protein within each pathway and the number of supporting evidences. This metric revealed that some pathway pairs (e.g., Wnt–β‑catenin with TGF‑β) exhibit particularly strong bidirectional communication, while others show more modest links.
The study further integrated tissue‑specific expression data from GTEx and cancer transcriptomes from TCGA. By mapping the presence of cross‑talk proteins onto these datasets, the authors identified tissue‑ and cancer‑specific cross‑talk clusters. For instance, liver tissue displays a pronounced Wnt–TGF‑β cross‑talk module, which aligns with known liver‑cancer signaling rewiring. Similar tissue‑restricted modules were observed in breast, lung, and colon cancers, suggesting that the same set of pathways can be repuned in a context‑dependent manner.
Leveraging the cross‑talk network, the authors performed a drug‑target discovery analysis. They defined “multi‑pathway tissue‑specific targets” as proteins that (i) participate in two or more signaling pathways, (ii) are expressed in a limited set of tissues or tumor types, and (iii) have supporting literature evidence. This filtering yielded 327 candidate proteins. Cross‑referencing with the FDA‑approved drug database showed that 42 of these candidates are already known drug targets (many of them anticancer agents), while the remaining 285 represent novel opportunities for network‑based drug repurposing or de‑novo drug design. Network centrality measures (betweenness, degree, clustering coefficient) further prioritized candidates that act as bottlenecks or hubs within the cross‑talk landscape.
Compared with established resources such as KEGG, Reactome, and Pathway Commons, SignaLink’s unique contribution lies in its uniform curation across species and pathways, enabling direct quantitative comparison of cross‑talks. The authors acknowledge that limiting the resource to eight pathways restricts the view of the full signaling universe, and that many predicted cross‑talk interactions lack experimental validation. Nonetheless, the database is openly accessible (http://SignaLink.org) and provides a valuable scaffold for systems‑biology investigations, precision‑medicine strategies, and hypothesis‑driven drug discovery. Future work will expand the pathway repertoire, incorporate additional model organisms, and integrate functional validation data to further enhance the utility of SignaLink.
Comments & Academic Discussion
Loading comments...
Leave a Comment