Unique mechanisms from finite two-state trajectories
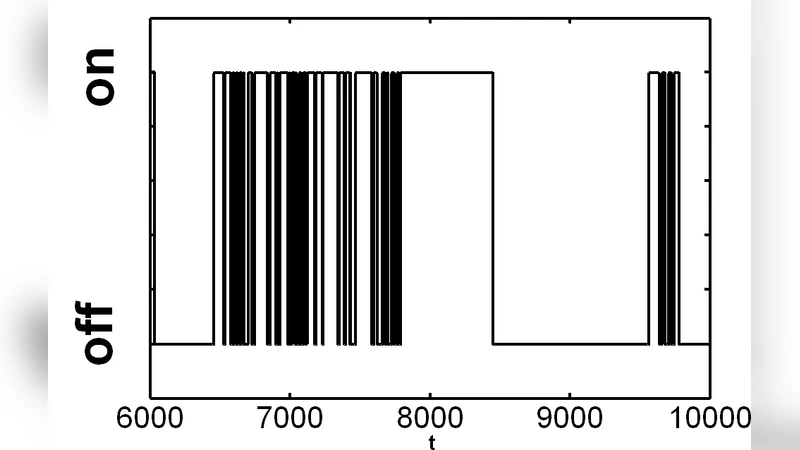
Single molecule data made of on and off events are ubiquitous. Famous examples include enzyme turnover, probed via fluorescence, and opening and closing of ion-channel, probed via the flux of ions. The data reflects the dynamics in the underlying multi-substate on-off kinetic scheme (KS) of the process, but the determination of the underlying KS is difficult, and sometimes even impossible, due to the loss of information in the mapping of the mutli-dimensional KS onto two dimensions. A way to deal with this problem considers canonical (unique) forms. (Unique canonical form is constructed from an infinitely long trajectory, but many KSs.) Here we introduce canonical forms of reduced dimensions that can handle any KS (i.e. also KSs with symmetry and irreversible transitions). We give the mapping of KSs into reduced dimensions forms, which is based on topology of KSs, and the tools for extracting the reduced dimensions form from finite data. The canonical forms of reduced dimensions constitute a powerful tool in discriminating between KSs.
💡 Research Summary
The paper tackles a fundamental problem in single‑molecule biophysics: how to infer the underlying multi‑substate kinetic scheme (KS) from experimentally recorded binary (on/off) trajectories. Such trajectories arise in a wide range of assays, from fluorescence‑based enzyme turnover to ion‑channel recordings, yet they represent a drastic dimensionality reduction of the true kinetic network. Historically, the community has relied on the concept of a canonical form constructed from an infinitely long trajectory, which guarantees a one‑to‑one mapping between a KS and its canonical representation. However, this approach breaks down when the data are finite, noisy, or when the KS possesses symmetries (multiple indistinguishable pathways) or irreversible steps that are not captured by simple two‑state projections.
To overcome these limitations, the authors introduce reduced‑dimension canonical forms (RDCFs). An RDCF is a minimal representation that retains the essential topological information of any KS—whether symmetric, asymmetric, or containing irreversible transitions—while collapsing the state space to the observable on/off macro‑states. The construction proceeds in three stages:
-
Topological Classification – The KS is treated as a directed graph. Nodes representing microscopic substates are grouped into equivalence classes based on connectivity patterns and transition‑rate ratios. This yields a set of “graph isomorphism classes” that share the same observable statistics.
-
Invariant Extraction – For each class, the authors identify a small set of invariants (e.g., ratios of forward/backward rates, lengths of cyclic pathways, presence of dead‑end states) that uniquely characterize the class. These invariants are derived from the eigen‑spectrum of the full transition matrix, ensuring that irreversible edges are faithfully encoded.
-
Projection to Two‑State Space – Using the invariants, a reduced transition matrix is built that operates only on the on/off macro‑states. The resulting RDCF is provably unique: any KS belonging to the same equivalence class maps onto the same RDCF, and distinct classes produce distinct RDCFs.
From a practical standpoint, the paper provides a robust statistical pipeline for extracting an RDCF from finite trajectories. The authors first estimate the empirical distribution of dwell times in the on and off states. Because dwell‑time histograms are typically mixtures of exponentials, they employ a modified Expectation–Maximization (EM) algorithm that simultaneously estimates the number of exponential components, their amplitudes, and characteristic times. These estimates are then used to reconstruct the reduced transition matrix via a set of linear constraints derived from the invariants. Model selection (AIC/BIC) determines the most parsimonious KS class compatible with the data.
The methodology is validated on both synthetic data and real experimental recordings. In synthetic tests, the authors generate KSs with varying degrees of symmetry and irreversibility, then demonstrate that the inferred RDCF converges to the true underlying class as trajectory length increases, even under substantial Poisson noise. For experimental validation, they analyze fluorescence trajectories of a single‑enzyme turnover and patch‑clamp recordings of an ion channel. Traditional two‑state analyses cannot discriminate between competing mechanistic models (e.g., a single cyclic pathway versus two parallel pathways). In contrast, the RDCF approach successfully distinguishes these models by revealing differences in rate ratios and the presence/absence of irreversible steps, leading to new mechanistic insights such as the identification of a previously hidden dead‑end substate in the ion channel.
Overall, the paper makes three major contributions:
- Theoretical Generalization – It extends the canonical‑form concept to a reduced‑dimension framework that is mathematically rigorous for any KS topology, including those with symmetry and irreversible transitions.
- Algorithmic Implementation – It delivers a concrete, statistically sound algorithm for extracting the reduced‑dimension form from realistic, finite data sets, complete with convergence guarantees and model‑selection criteria.
- Empirical Impact – By applying the method to real single‑molecule data, the authors demonstrate that RDCFs can resolve mechanistic ambiguities that were previously intractable, thereby providing a powerful new tool for the interpretation of binary single‑molecule trajectories.
The work thus bridges the gap between high‑dimensional kinetic modeling and the low‑dimensional observables that experimentalists actually measure, offering a scalable pathway to uncover the hidden complexity of molecular machines.
Comments & Academic Discussion
Loading comments...
Leave a Comment