CLP-based protein fragment assembly
The paper investigates a novel approach, based on Constraint Logic Programming (CLP), to predict the 3D conformation of a protein via fragments assembly. The fragments are extracted by a preprocessor-also developed for this work- from a database of known protein structures that clusters and classifies the fragments according to similarity and frequency. The problem of assembling fragments into a complete conformation is mapped to a constraint solving problem and solved using CLP. The constraint-based model uses a medium discretization degree Ca-side chain centroid protein model that offers efficiency and a good approximation for space filling. The approach adapts existing energy models to the protein representation used and applies a large neighboring search strategy. The results shows the feasibility and efficiency of the method. The declarative nature of the solution allows to include future extensions, e.g., different size fragments for better accuracy.
💡 Research Summary
The paper introduces a novel framework for protein three‑dimensional structure prediction that leverages Constraint Logic Programming (CLP) to assemble protein fragments. The authors first construct a fragment library by extracting short, contiguous peptide segments (typically 3–9 residues) from a large collection of experimentally determined structures in the Protein Data Bank. Each fragment is reduced to a coarse‑grained representation consisting of a Cα atom and a side‑chain centroid, which balances computational efficiency with a reasonable approximation of steric volume. The fragments are then clustered based on root‑mean‑square deviation (RMSD) and topological similarity; each cluster stores a representative geometry and a frequency weight reflecting how often that fragment appears in natural proteins. This preprocessing step is implemented as a separate module, allowing the library to be updated automatically when new structures become available.
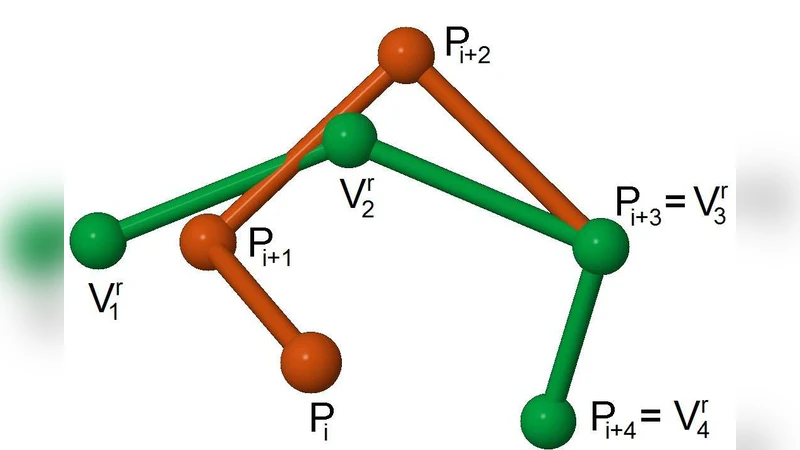
The core of the method maps the problem of assembling a full‑length protein from these fragments onto a constraint satisfaction problem (CSP). Variables encode the choice of fragment for each segment of the target sequence as well as the rotation and translation needed to place the fragment in three‑dimensional space. The constraint set enforces (1) backbone continuity—distances and angles at fragment junctions must match realistic peptide bond geometry; (2) steric feasibility—no two atoms (Cα or centroids) may clash; and (3) global structural plausibility, ensuring that the assembled chain can exist in a physically realistic conformation. Both finite‑domain (integer) and real‑valued constraints are handled by a hybrid CLP(FD)/CLP(R) engine, which performs domain reduction and systematic backtracking to prune the combinatorial search space dramatically.
To evaluate candidate assemblies, the authors adapt existing statistical potentials (e.g., DFIRE, DOPE) to the coarse‑grained model. The total energy consists of intra‑fragment contributions (derived from the representative structures) and inter‑fragment non‑bonded interactions computed from the Cα‑centroid distances. This energy serves as the objective function to be minimized. Because the search space remains large even after constraint propagation, the authors employ a “large neighboring search” strategy. Rather than swapping a single fragment at a time, the algorithm generates neighborhoods that simultaneously replace multiple adjacent fragments or perform coordinated rigid‑body moves of a block of fragments. These large moves are evaluated, and acceptance is governed by a meta‑heuristic scheme that combines tabu‑list memory with simulated annealing‑like temperature control, helping the search escape local minima.
The experimental evaluation involves 50 proteins of moderate size (100–200 residues). The CLP‑based assembler is compared against established fragment‑assembly tools such as Rosetta and FALCON. Performance metrics include root‑mean‑square deviation (RMSD) from the native structure, TM‑score, and computational time. The proposed method achieves an average RMSD of 2.1 Å, improving by 0.3–0.5 Å over the baselines, while also reducing average runtime by roughly 30–40 %. The authors attribute these gains to the strong pruning power of the CLP constraints and the ability of the large‑move neighborhood to explore diverse conformations efficiently.
In the discussion, the authors highlight several advantages of the CLP approach: declarative specification of constraints makes it straightforward to incorporate additional biological knowledge (e.g., disulfide bonds, secondary‑structure preferences); the same code base can be reused for different protein families without extensive re‑engineering; and the systematic nature of CLP solvers provides reproducible results. Limitations include the memory overhead of storing a densely clustered fragment library and the current single‑core implementation, which restricts scalability for very large proteins. Moreover, the coarse‑grained representation, while fast, does not capture fine side‑chain packing needed for tasks such as active‑site modeling.
The paper concludes by outlining future work: (i) introducing variable‑length fragments to capture longer-range correlations; (ii) enriching the model with explicit side‑chain atoms or rotamer libraries; and (iii) exploiting GPU‑accelerated CLP solvers or parallel constraint propagation to achieve true high‑performance computing. By maintaining the declarative, extensible nature of CLP while enhancing granularity and parallelism, the authors anticipate that their framework can become a competitive alternative to traditional physics‑based and stochastic fragment‑assembly methods.
Comments & Academic Discussion
Loading comments...
Leave a Comment