Dominant folding pathways of a peptide chain, from ab-initio quantum-mechanical simulations
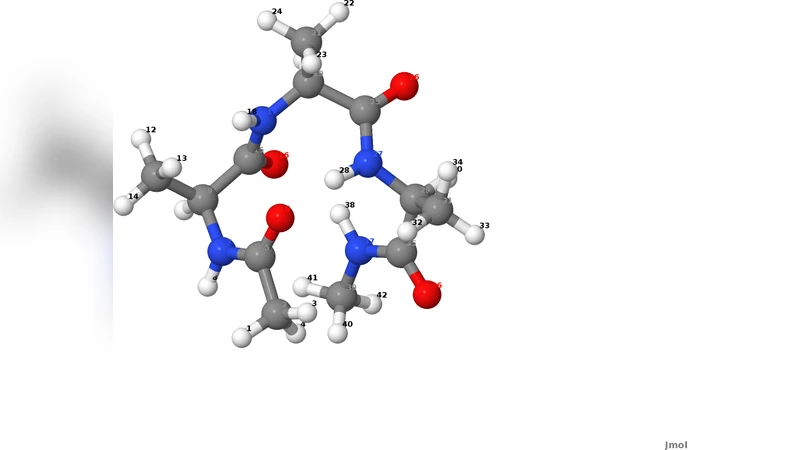
Using the Dominant Reaction Pathways method, we perform an ab-initio quantum-mechanical simulation of a conformational transition of a peptide chain. The method we propose makes it possible to investigate the out-of-equilibrium dynamics of these systems, without resorting to an empirical representation of the molecular force field. It also allows to study rare transitions involving rearrangements in the electronic structure. By comparing the results of the ab-initio simulation with those obtained employing a standard force field, we discuss its capability to describe the non-equilibrium dynamics of conformational transitions.
💡 Research Summary
The paper presents a novel implementation of the Dominant Reaction Pathways (DRP) method that incorporates ab‑initio quantum‑mechanical forces, thereby enabling the study of peptide folding transitions without relying on empirical force fields. Traditional molecular dynamics (MD) simulations of protein folding use classical force fields, which are parametrized to reproduce equilibrium properties but cannot capture rare events that involve significant electronic rearrangements. The authors address this limitation by embedding density‑functional theory (DFT) calculations directly into the DRP framework, allowing the simultaneous evolution of nuclear coordinates and electronic density along the most probable transition pathway.
The methodological core consists of three steps. First, the stochastic Langevin dynamics of the peptide are expressed as a path‑integral, leading to an action functional S
Comments & Academic Discussion
Loading comments...
Leave a Comment