A population-based microbial oscillator
Genetic oscillators are a major theme of interest in the emerging field of synthetic biology. Until recently, most work has been carried out using intra-cellular oscillators, but this approach restricts the broader applicability of such systems. Motivated by a desire to develop large-scale, spatially-distributed cell-based computational systems, we present an initial design for a population-level oscillator which uses three different bacterial strains. Our system is based on the client-server model familiar to computer science, and uses quorum sensing for communication between nodes. We present the results of extensive in silico simulation tests, which confirm that our design is both feasible and robust.
💡 Research Summary
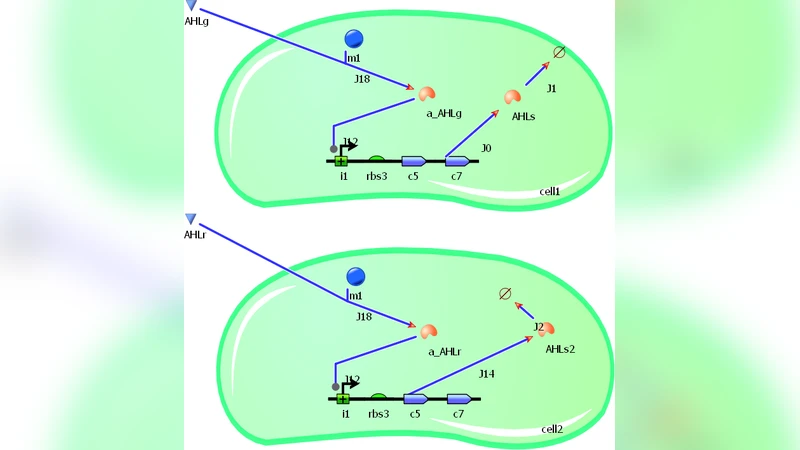
The paper introduces a novel design for a population‑level genetic oscillator that operates across three engineered bacterial strains, moving beyond the traditional intracellular oscillators that have dominated synthetic biology. Drawing inspiration from the client‑server architecture in computer science, the authors assign distinct functional roles to each strain: a “server” strain that simultaneously synthesizes two quorum‑sensing autoinducers (AHL‑1 and AHL‑2), and two “client” strains (Client‑A and Client‑B) that respond reciprocally to these signals. Server‑derived AHL‑1 triggers the expression of a repressor in Client‑A while simultaneously suppressing genes activated by AHL‑2; conversely, AHL‑2 activates a repressor in Client‑B and inhibits genes responsive to AHL‑1. This cross‑inhibition creates a feedback loop that forces the entire bacterial population to cycle through defined transcriptional states in a synchronized manner.
To evaluate feasibility, the authors constructed deterministic ordinary differential equation (ODE) models and stochastic Gillespie simulations of the coupled reaction network. Parameter sweeps examined the impact of synthesis rates, diffusion coefficients, degradation constants, and initial strain ratios. The results demonstrate robust oscillatory behavior across a wide range of conditions: variations of up to 10 % in initial strain composition, temperature fluctuations of ±20 %, and the presence or absence of AHL‑degrading enzymes all produced only minor changes in period and amplitude. Sensitivity analysis identified the server’s AHL production capacity as the critical threshold; below this level oscillations collapse, a limitation that can be mitigated by employing stronger promoters or higher‑efficiency synthases.
Stochastic simulations further revealed that intrinsic noise inherent to low‑cell‑number environments does not destabilize the collective rhythm. The mutual feedback between the three strains effectively averages out fluctuations, preserving a consistent period and amplitude at the population level. This noise‑tolerance is a key advantage over single‑cell oscillators, which often require additional buffering mechanisms.
The discussion highlights several strategic benefits of the proposed system. First, the spatially distributed architecture enables scaling to large, heterogeneous environments without the need for precise intracellular timing mechanisms. Second, the use of quorum sensing as a communication medium provides a chemically simple yet versatile channel for information exchange, allowing the implementation of more complex logical operations in future multi‑layered bio‑computing networks. Third, the design exhibits strong resilience to environmental perturbations, making it a promising platform for real‑world applications such as synchronized drug delivery, biosensing arrays, or programmable bioproduction.
The authors acknowledge practical challenges that must be addressed before experimental realization. These include potential cross‑talk among AHL molecules, differential growth rates among the engineered strains, and diffusion limitations in bulk cultures. They propose solutions such as orthogonal quorum‑sensing systems, growth‑rate balancing via metabolic engineering, and the use of microfluidic devices to control spatial distribution and signal gradients.
In conclusion, the study provides a comprehensive computational validation of a client‑server‑based, population‑wide microbial oscillator. By demonstrating that synchronized oscillations can emerge from simple, distributed quorum‑sensing interactions, the work opens a new avenue for constructing large‑scale, spatially distributed synthetic biological circuits that mimic the modularity and robustness of conventional computing architectures.
Comments & Academic Discussion
Loading comments...
Leave a Comment